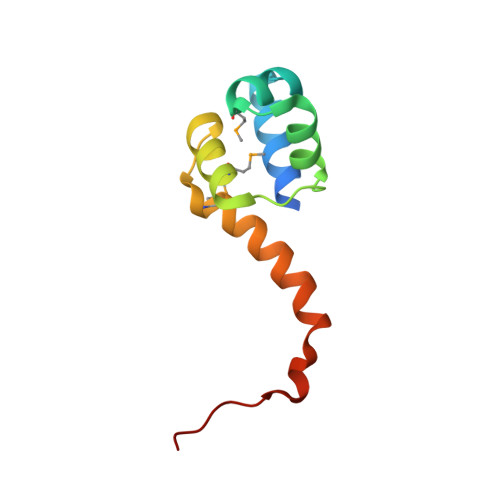
Crystal structure of the bacterial antitoxin HigA from Escherichia coli.
Arbing, M.A., Abashidze, M., Hurley, J.M., Zhao, L., Janjua, H., Cunningham, K., Ma, L.C., Xiao, R., Liu, J., Baran, M.C., Acton, T.B., Rost, B., Inouye, M., Woychik, N.A., Montelione, G.T., Hunt, J.F.To be published.