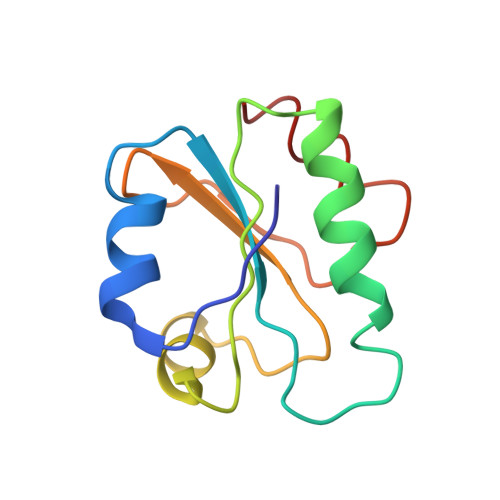
NMR solution structure of the reduced form of thioredoxin 1 from Sacharomyces cerevisiae
Pinheiro, A.S., Amorim, G.C., Netto, L.E., Almeida, F.C., Valente, A.P.(2008) Proteins 70: 584-587
- PubMed: 17932921 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21693
- Primary Citation Related Structures:
2I9H - Centro Nacional de Ressonância Magnética Nucelar Jiri Jonas, Instituo de Bioquímica Médica, Universidade Federal do Rio de Janeiro, Rio de Janeiro-RJ, Brasil.
Organizational Affiliation: