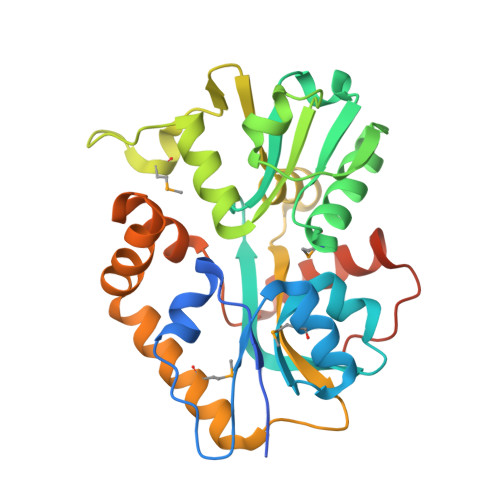
X-ray structures of two proteins belonging to Pfam DUF178 revealed unexpected structural similarity to the DUF191 Pfam family.
Tyagi, R., Burley, S.K., Swaminathan, S.(2007) BMC Struct Biol 7: 62-62
- PubMed: 17908300 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-7-62
- Primary Citation Related Structures:
2I6E - PubMed Abstract:
Pfam is a comprehensive collection of protein domains and families, with a range of well-established information including genome annotation. Pfam has two large series of functionally uncharacterized families, known as Domains of Unknown Function (DUFs) and Uncharacterized Protein Families (UPFs). Crystal structures of two proteins from Deinococcus radiodurans and Streptomyces coelicolor belonging to Pfam protein family DUF178 (ID: PF02621) have been determined using Selenium-Single-wavelength Anomalous Dispersion (Se-SAD). Based on the structure, we have identified the putative function for this family of protein. Unexpectedly, we found that DUF178 Pfam is remarkably similar to Pfam family DUF191 suggesting that the sequence-based classification alone may not be sufficient to classify proteins into Pfam families.
- Biology Department, Brookhaven National Laboratory, Upton, New York 11973, USA. swami@bnl.gov
Organizational Affiliation: