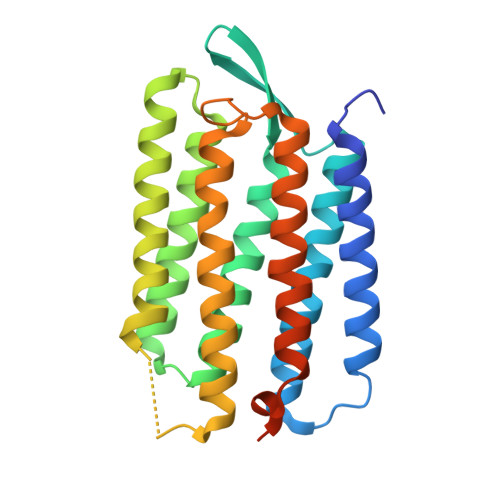
Propagating Structural Perturbation Inside Bacteriorhodopsin: Crystal Structures of the M State and the D96A and T46V Mutants.
Lanyi, J.K., Schobert, B.(2006) Biochemistry 45: 12003-12010
- PubMed: 17002299 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi061310i
- Primary Citation Related Structures:
2I1X, 2I20, 2I21 - PubMed Abstract:
The X-ray diffraction structure of the non-illuminated D96A bacteriorhodopsin mutant reveals structural changes as far away as 15 A from residue 96, at the retinal, Trp-182, Ala-215, and waters 501, 402, and 401. The Asp-to-Ala side-chain replacement breaks its hydrogen bond with Thr-46, and the resulting separation of the cytoplasmic ends of helices B and C is communicated to the retinal region through a chain of covalent and hydrogen bonds. The unexpected long-range consequences of the D96A mutation include breaking the hydrogen bond between O of Ala-215 and water 501 and the formation of a new hydrogen bond between water molecules 401 and 402 in the extracellular region. Because in the T46V mutant a new water molecule appears at Asp-96 and its hydrogen-bond to Ile-45 replaces Thr-46 as its link to helix B, the separation of helices B and C is smaller than that in D96A, and there are no atomic displacements elsewhere in the protein. Propagation of conformational changes along the chain between the retinal and Thr-46 had been observed earlier in the crystal structures of the D96N and E204Q mutants but in the trapped M state. Consistent with the perturbation of the retinal region in D96A, little change of the Thr-46 region occurs between the non-illuminated and M states of this mutant. It appears that a local perturbation can propagate along a track in both directions between the retinal and the Asp-96/Thr-46 pair, either from photoisomerization of the retinal in the wild-type protein in one case or from the D96A mutation in the other.
- Department of Physiology and Biophysics, University of California, Irvine, California 92697, USA. jlanyi@orion.oac.uci.edu
Organizational Affiliation: