Stereochemistry Modulates the Stability of Reduced Interstrand Cross-Links Arising from R- and S-alpha-CH(3)-gamma-OH-1,N(2)-Propano-2'-deoxyguanosine in the 5'-CpG-3' DNA Sequence
Cho, Y.-J., Kozekov, I.D., Harris, T.M., Rizzo, C.J., Stone, M.P.(2007) Biochemistry 46: 2608-2621
- PubMed: 17305317 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi061381h
- Primary Citation Related Structures:
2HMD, 2HMR - PubMed Abstract:
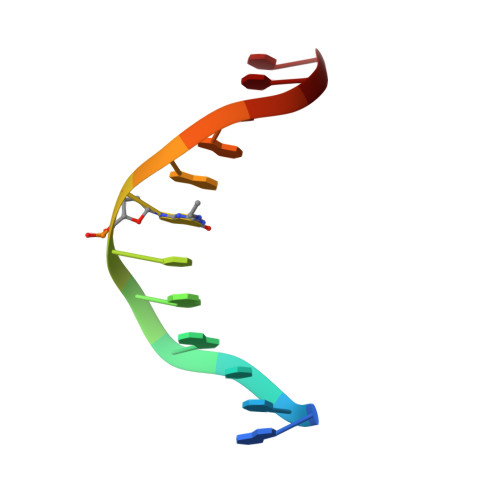
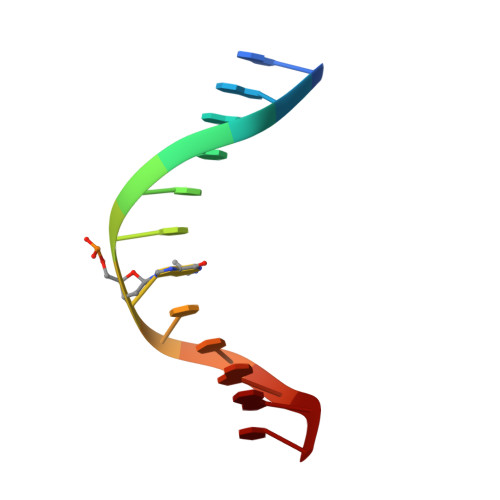
The solution structures of 5'-Cp-N2-dG-3'-R-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' and 5'-Cp-N2-dG-3'-S-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' interstrand DNA cross-links in the 5'-CpG-3' sequence were determined by NMR spectroscopy. These were utilized as chemically stable surrogates for the corresponding carbinolamine interstrand cross-links arising from the crotonaldehyde- and acetaldehyde-derived R- and S-alpha-CH3-gamma-OH-1,N2-propanodeoxyguanosine adducts. The results provide an explanation for the observation that interstrand cross-link formation in the 5'-CpG-3' sequence by the R- and S-alpha-CH3-gamma-OH-1,N2-propanodeoxyguanosine adducts is dependent upon stereochemistry, favoring the R-alpha-CH3-gamma-OH-1,N2-propanodeoxyguanosine adduct [Kozekov, I. D., Nechev, L. V., Moseley, M. S., Harris, C. M., Rizzo, C. J., Stone, M. P., and Harris, T. M. (2003) J. Am. Chem. Soc. 125, 50-61]. Molecular dynamics calculations, restrained by NOE-based distances and empirical restraints, revealed that both the 5'-Cp-N2-dG-3'-R-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' and 5'-Cp-N2-dG-3'-S-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' cross-links were located in the minor groove and retained Watson-Crick hydrogen bonds at the tandem cross-linked C.G base pairs. However, for the 5'-Cp-N2-dG-3'-R-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' cross-link, the (alpha)-CH3 group was positioned in the center of the minor groove, whereas for the 5'-Cp-N2-dG-3'-S-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' cross-link, the (alpha)-CH3 group was positioned in the 3' direction, showing steric interference with the DNA helix. The 5'-Cp-N2-dG-3'-S-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' cross-link exhibited a lower thermal stability as evidenced by NMR spectroscopy as a function of temperature. The two cross-links also exhibited apparent differences in the conformation of the interstrand three-carbon cross-link, which may also contribute to the lower apparent thermodynamic stability of the 5'-Cp-N2-dG-3'-S-(alpha)-CH3-propyl-5'-Cp-N2-dG-3' cross-link.
- Department of Chemistry, Center in Molecular Toxicology, Vanderbilt-Ingram Cancer Center, Vanderbilt University, Nashville, Tennessee 37235, USA.
Organizational Affiliation: