Mechanism of Intracellular Block of the KcsA K(+) Channel by Tetrabutylammonium: Insights from X-ray Crystallography, Electrophysiology and Replica-exchange Molecular Dynamics Simulations.
Faraldo-Gomez, J.D., Kutluay, E., Jogini, V., Zhao, Y., Heginbotham, L., Roux, B.(2007) J Mol Biology 365: 649-662
- PubMed: 17070844 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.09.069
- Primary Citation Related Structures:
2HJF - PubMed Abstract:
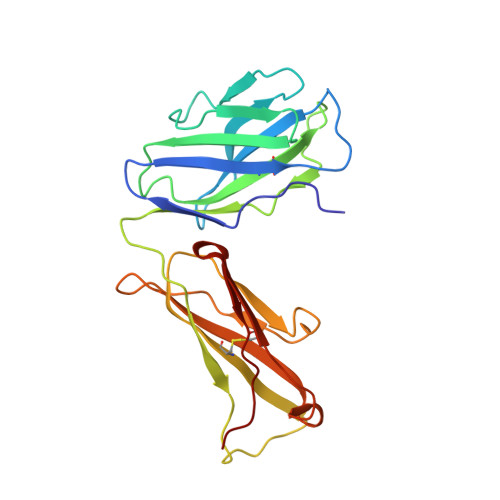
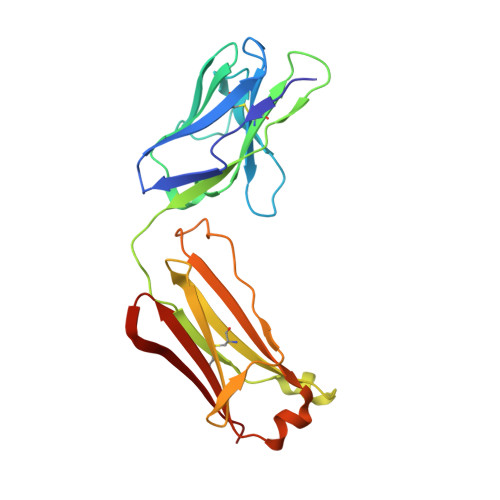
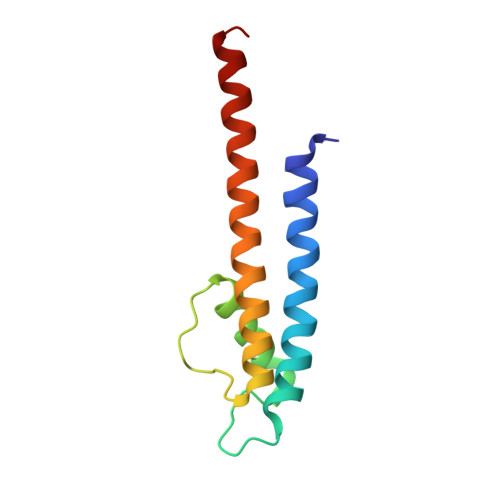
The mechanism of intracellular blockade of the KcsA potassium channel by tetrabutylammonium (TBA) is investigated through functional, structural and computational studies. Using planar-membrane electrophysiological recordings, we characterize the binding kinetics as well as the dependence on the transmembrane voltage and the concentration of the blocker. It is found that the apparent affinity of the complex is significantly greater than that of any of the eukaryotic K(+) channels studied previously, and that the off-rate increases with the applied transmembrane voltage. In addition, we report a crystal structure of the KcsA-TBA complex at 2.9 A resolution, with TBA bound inside the large hydrophobic cavity located at the center of the channel, consistent with the results of previous functional and structural studies. Of particular interest is the observation that the presence of TBA has a negligible effect on the channel structure and on the position of the potassium ions occupying the selectivity filter. Inspection of the electron density corresponding to TBA suggests that the ligand may adopt more than one conformation in the complex, though the moderate resolution of the data precludes a definitive interpretation on the basis of the crystallographic refinement methods alone. To provide a rationale for these observations, we carry out an extensive conformational sampling of an atomic model of TBA bound in the central cavity of KcsA, using the Hamiltonian replica-exchange molecular dynamics simulation method. Comparison of the simulated and experimental density maps indicates that the latter does reflect at least two distinct binding orientations of TBA. The simulations show also that the relative population of these binding modes is dependent on the ion configuration occupying the selectivity filter, thus providing a clue to the nature of the voltage-dependence of the binding kinetics.
- Institute for Molecular Pediatric Sciences and Department of Biochemistry and Molecular Biology, Gordon Center for Integrative Sciences, University of Chicago, 929 East 57th Street, Chicago, IL 60637, USA.
Organizational Affiliation: