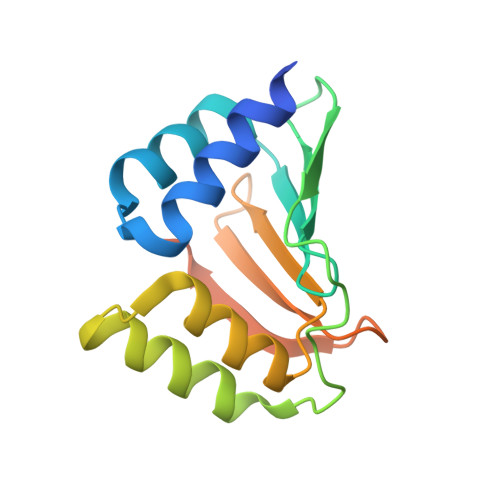
Solution NMR structure of the ydfO protein from Escherichia coli. Northeast Structural Genomics target ER251.
Rossi, P., Cort, J.R., Ho, C.K., Janjua, H., Cunningham, K., Ma, L.-C., Xiao, R., Liu, J., Baran, M., Swapna, G.V.T., Acton, T.B., Rost, B., Kennedy, M.A., Montelione, G.T.To be published.