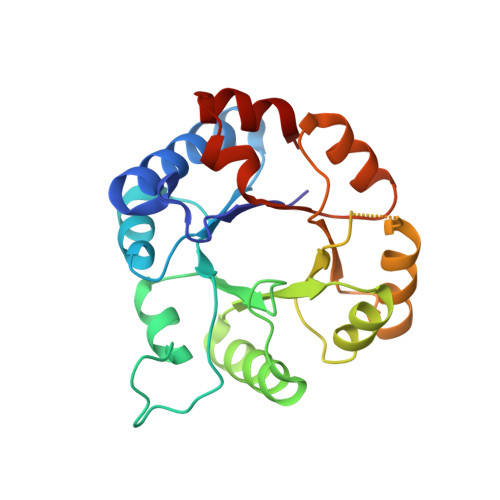
Structure of triosephosphate isomerase (TIM) from Methanocaldococcus jannaschii
Gayathri, P., Banerjee, M., Vijayalakshmi, A., Azeez, S., Balaram, H., Balaram, P., Murthy, M.R.N.(2007) Acta Crystallogr D Biol Crystallogr 63: 206-220
- PubMed: 17242514 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444906046488
- Primary Citation Related Structures:
2H6R - PubMed Abstract:
The crystal structure of a recombinant triosephosphate isomerase (TIM) from the archaeabacterium Methanocaldococcus jannaschii has been determined at a resolution of 2.3 A using X-ray diffraction data from a tetartohedrally twinned crystal. M. jannaschii TIM (MjTIM) is tetrameric, as suggested by solution studies and from the crystal structure, as is the case for two other structurally characterized archaeal TIMs. The archaeabacterial TIMs are shorter compared with the dimeric TIMs; the insertions in the dimeric TIMs occur in the vicinity of the tetramer interface, resulting in a hindrance to tetramerization in the dimeric TIMs. The charge distribution on the surface of the archaeal TIMs also facilitates tetramerization. Analysis of the barrel interactions in TIMs suggests that these interactions are unlikely to account for the thermal stability of the archaeal TIMs. A novelty of the unliganded structure of MjTIM is the complete absence of electron density for the loop 6 residues. The disorder of this loop could be ascribed to a missing salt bridge between residues at the N- and C-terminal ends of the loop in MjTIM.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.
Organizational Affiliation: