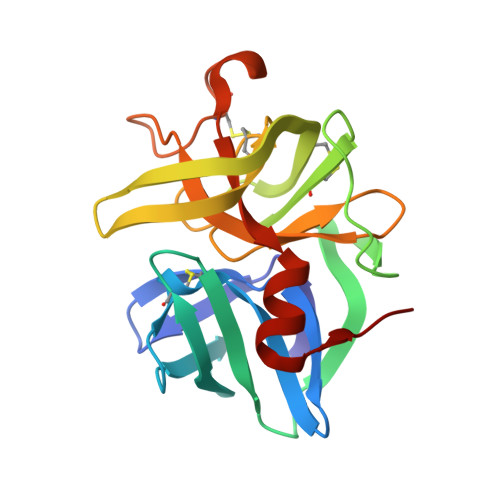
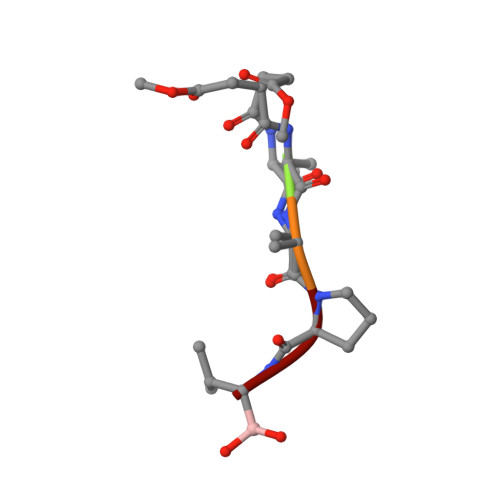
Subangstrom crystallography reveals that short ionic hydrogen bonds, and not a His-Asp low-barrier hydrogen bond, stabilize the transition state in serine protease catalysis
Fuhrmann, C.N., Daugherty, M.D., Agard, D.A.(2006) J Am Chem Soc 128: 9086-9102
- PubMed: 16834383 Search on PubMed
- DOI: https://doi.org/10.1021/ja057721o
- Primary Citation Related Structures:
2H5C, 2H5D - PubMed Abstract:
To address questions regarding the mechanism of serine protease catalysis, we have solved two X-ray crystal structures of alpha-lytic protease (alphaLP) that mimic aspects of the transition states: alphaLP at pH 5 (0.82 A resolution) and alphaLP bound to the peptidyl boronic acid inhibitor, MeOSuc-Ala-Ala-Pro-boroVal (0.90 A resolution). Based on these structures, there is no evidence of, or requirement for, histidine-flipping during the acylation step of the reaction. Rather, our data suggests that upon protonation of His57, Ser195 undergoes a conformational change that destabilizes the His57-Ser195 hydrogen bond, preventing the back-reaction. In both structures the His57-Asp102 hydrogen bond in the catalytic triad is a normal ionic hydrogen bond, and not a low-barrier hydrogen bond (LBHB) as previously hypothesized. We propose that the enzyme has evolved a network of relatively short hydrogen bonds that collectively stabilize the transition states. In particular, a short ionic hydrogen bond (SIHB) between His57 Nepsilon2 and the substrate's leaving group may promote forward progression of the TI1-to-acylenzyme reaction. We provide experimental evidence that refutes use of either a short donor-acceptor distance or a downfield 1H chemical shift as sole indicators of a LBHB.
- Howard Hughes Medical Institute and Department of Biochemistry and Biophysics, University of California, San Francisco, San Francisco, California 94143-2240, USA.
Organizational Affiliation: