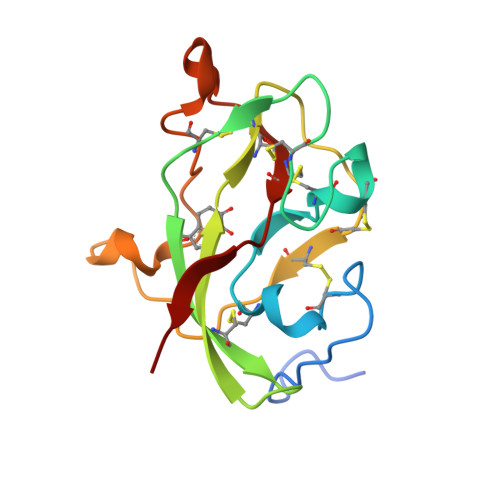
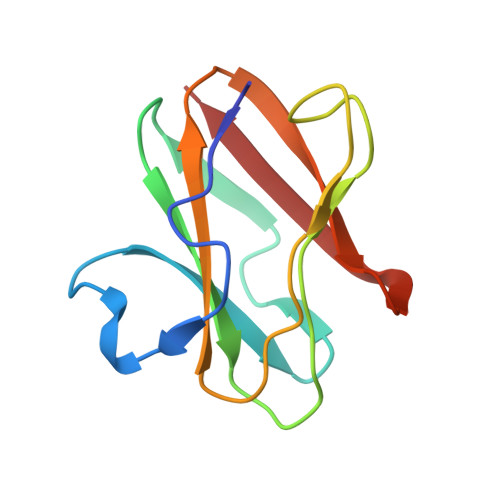
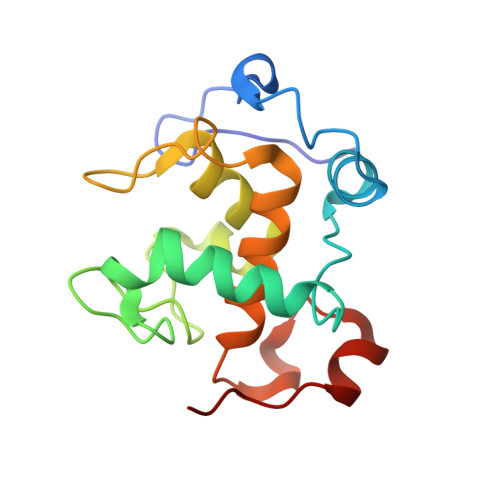
Structural comparison of the oxidized ternary electron transfer complex of methylamine dehydrogenase, amicyanin and cytochrome c551i from Paracoccus denitrificans with the substrate-reduced, copper free complex at 1.9 A resolution.
Chen, Z., Durley, R., Davidson, V.L., Mathews, F.S.To be published.