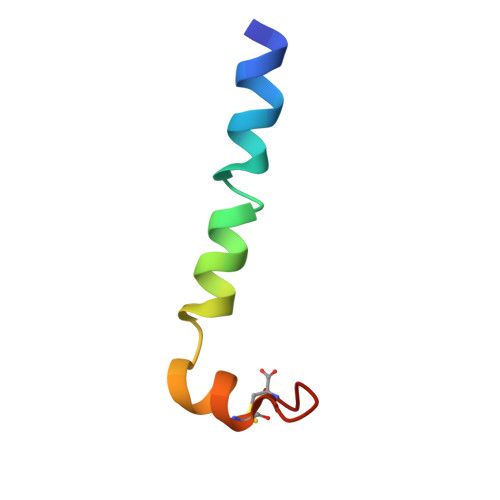
Solution structure and membrane interaction mode of an antimicrobial peptide gaegurin 4
Chi, S.-W., Kim, J.-S., Kim, D.-H., Lee, S.-H., Park, Y.-H., Han, K.-H.(2007) Biochem Biophys Res Commun 352: 592-597
- PubMed: 17141187 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2006.11.064
- Primary Citation Related Structures:
2G9L - PubMed Abstract:
We have applied NMR spectroscopy to determine the high-resolution structure of gaegurin 4, a 37-residue antimicrobial peptide from Rana rugosa, under varying hydrophobic conditions. Even in 100% H2O, gaegurin 4 contains a nascent turn near its C-terminal Rana box. Under a more hydrophobic condition it forms two amphipathic helices, one long encompassing residues 2-23 and the other consisting of residues 25-34, similar to what has been observed in cecropin A. Functional implication of the helix-breaking kink at Gly24 in gaegurin 4 was investigated by preparing several analogs. Based upon the current and previous results, we propose a novel seaanemone-like ion pore-forming model for gaegurin 4.
- Molecular Cancer Research Center, Division of Molecular Therapeutics, KRIBB, Daejeon 305-806, Republic of Korea.
Organizational Affiliation: