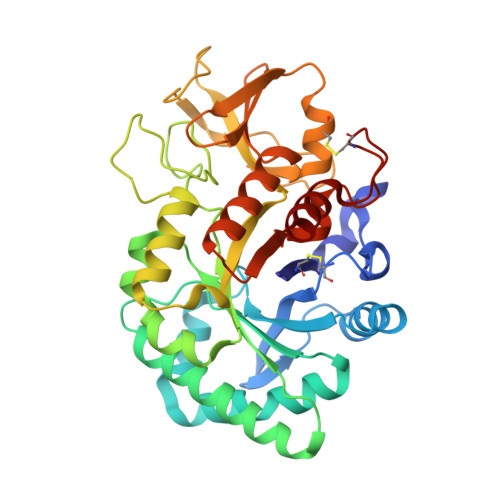
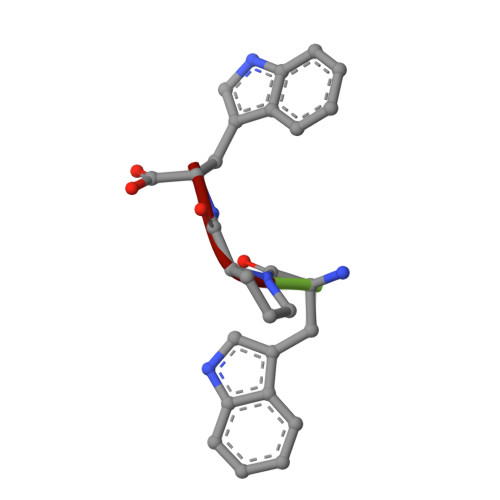
Crystal structure of the ternary complex of signalling protein from sheep (SPS-40) with trimer and designed peptide at 2.5A resolution
Ethayathulla, A.S., Srivastava, D.B., Kumar, J., Somvanshi, R.K., Bhushan, A., Sharma, S., Dey, S., Singh, T.P.To be published.