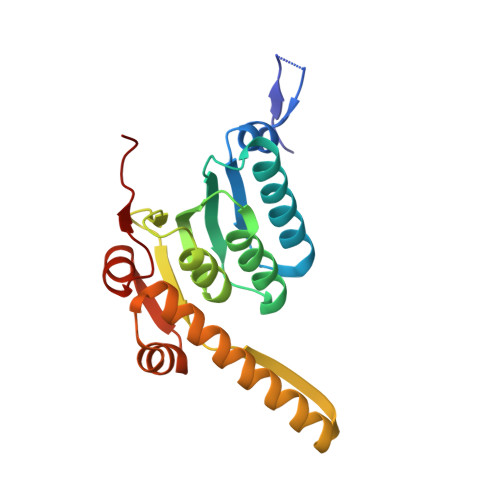
Crystal structure at 1.9A of E. coli ClpP with a peptide covalently bound at the active site.
Szyk, A., Maurizi, M.R.(2006) J Struct Biol 156: 165-174
- PubMed: 16682229 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2006.03.013
- Primary Citation Related Structures:
2FZS - PubMed Abstract:
ClpP, the proteolytic component of the ATP-dependent ClpAP and ClpXP chaperone/protease complexes, has 14 identical subunits organized in two stacked heptameric rings. The active sites are in an interior aqueous chamber accessible through axial channels. We have determined a 1.9 A crystal structure of Escherichia coli ClpP with benzyloxycarbonyl-leucyltyrosine chloromethyl ketone (Z-LY-CMK) bound at each active site. The complex mimics a tetrahedral intermediate during peptide cleavage, with the inhibitor covalently linked to the active site residues, Ser97 and His122. Binding is further stabilized by six hydrogen bonds between backbone atoms of the peptide and ClpP as well as by hydrophobic binding of the phenolic ring of tyrosine in the S1 pocket. The peptide portion of Z-LY-CMK displaces three water molecules in the native enzyme resulting in little change in the conformation of the peptide binding groove. The heptameric rings of ClpP-CMK are slightly more compact than in native ClpP, but overall structural changes were minimal (rmsd approximately 0.5 A). The side chain of Ser97 is rotated approximately 90 degrees in forming the covalent adduct with Z-LY-CMK, indicating that rearrangement of the active site residues to a active configuration occurs upon substrate binding. The N-terminal peptide of ClpP-CMK is stabilized in a beta-hairpin conformation with the proximal N-terminal residues lining the axial channel and the loop extending beyond the apical surface of the heptameric ring. The lack of major substrate-induced conformational changes suggests that changes in ClpP structure needed to facilitate substrate entry or product release must be limited to rigid body motions affecting subunit packing or contacts between ClpP rings.
- Laboratory of Cell Biology, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: