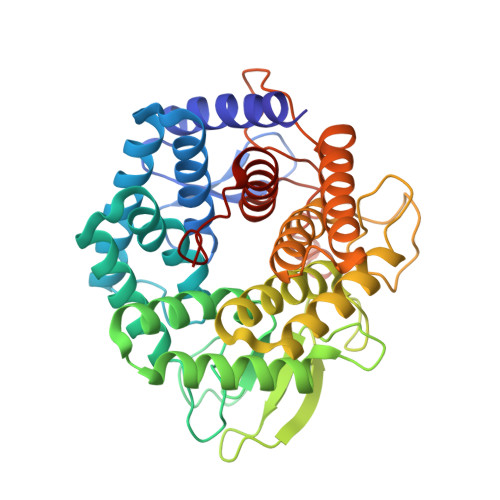
Substrate recognition by unsaturated glucuronyl hydrolase from Bacillus sp. GL1
Itoh, T., Hashimoto, W., Mikami, B., Murata, K.(2006) Biochem Biophys Res Commun 344: 253-262
- PubMed: 16630576 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2006.03.141
- Primary Citation Related Structures:
2FUZ, 2FV0, 2FV1 - PubMed Abstract:
Bacterial unsaturated glucuronyl hydrolases (UGLs) together with polysaccharide lyases are responsible for the complete depolymerization of mammalian extracellular matrix glycosaminoglycans. UGL acts on various oligosaccharides containing unsaturated glucuronic acid (DeltaGlcA) at the nonreducing terminus and releases DeltaGlcA through hydrolysis. In this study, we demonstrate the substrate recognition mechanism of the UGL of Bacillus sp. GL1 by determining the X-ray crystallographic structure of its substrate-enzyme complexes. The tetrasaccharide-enzyme complex demonstrated that at least four subsites are present in the active pocket. Although several amino acid residues are crucial for substrate binding, the enzyme strongly recognizes DeltaGlcA at subsite -1 through the formation of hydrogen bonds and stacking interactions, and prefers N-acetyl-d-galactosamine and glucose rather than N-acetyl-d-glucosamine as a residue accommodated in subsite +1, due to the steric hindrance.
- Division of Agronomy and Horticultural Science, Graduate School of Agriculture, Kyoto University, Gokasho, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: