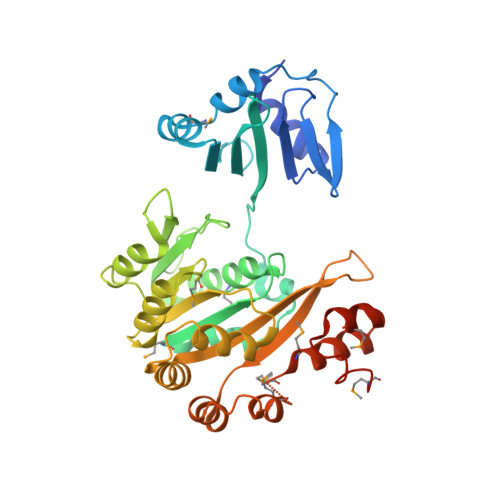
Crystal structures of the pilus retraction motor PilT suggest large domain movements and subunit cooperation drive motility.
Satyshur, K.A., Worzalla, G.A., Meyer, L.S., Heiniger, E.K., Aukema, K.G., Misic, A.M., Forest, K.T.(2007) Structure 15: 363-376
- PubMed: 17355871 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2007.01.018
- Primary Citation Related Structures:
2EWV, 2EWW, 2EYU, 2GSZ - PubMed Abstract:
PilT is a hexameric ATPase required for bacterial type IV pilus retraction and surface motility. Crystal structures of ADP- and ATP-bound Aquifex aeolicus PilT at 2.8 and 3.2 A resolution show N-terminal PAS-like and C-terminal RecA-like ATPase domains followed by a set of short C-terminal helices. The hexamer is formed by extensive polar subunit interactions between the ATPase core of one monomer and the N-terminal domain of the next. An additional structure captures a nonsymmetric PilT hexamer in which approach of invariant arginines from two subunits to the bound nucleotide forms an enzymatically competent active site. A panel of pilT mutations highlights the importance of the arginines, the PAS-like domain, the polar subunit interface, and the C-terminal helices for retraction. We present a model for ATP binding leading to dramatic PilT domain motions, engagement of the arginine wire, and subunit communication in this hexameric motor. Our conclusions apply to the entire type II/IV secretion ATPase family.
- Department of Bacteriology, University of Wisconsin-Madison, Madison, WI 53706, USA.
Organizational Affiliation: