Structure of circullin B and implications for antimicrobial activity of the cyclotides
Koltay, A., Daly, N.L., Gustafson, K.R., Craik, D.J.(2005) Int J Pept Protein Res 11: 99-106
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2005) Int J Pept Protein Res 11: 99-106
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Circulin B | 31 | Chassalia parviflora | Mutation(s): 0 | 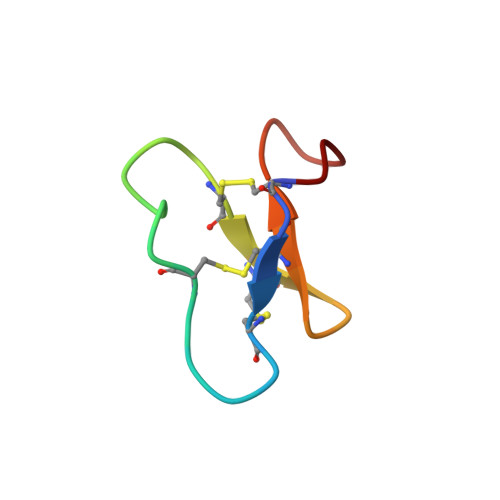 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P56879 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||