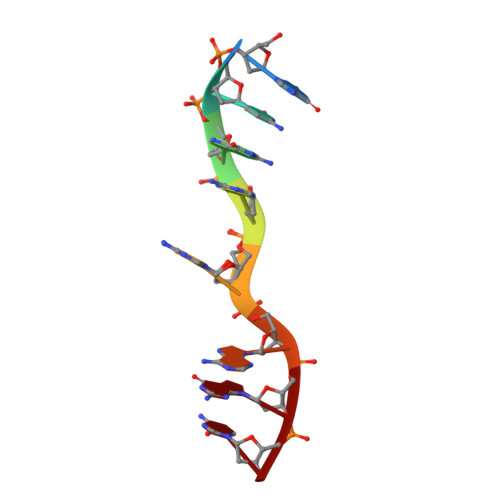
DNA octaplex formation with an I-motif of water-mediated A-quartets: reinterpretation of the crystal structure of d(GCGAAAGC)
Sato, Y., Mitomi, K., Sunami, T., Kondo, J., Takenaka, A.(2006) J Biochem 140: 759-762
- PubMed: 17062599 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvj213
- Primary Citation Related Structures:
2DZ7 - PubMed Abstract:
The crystal structure of the tetragonal form of d(gcGAAAgc) has been revised and reasonably refined including the disordered residues. The two DNA strands form a base-intercalated duplex, and the four duplexes are assembled according to the crystallographic 222 symmetry to form an octaplex. In the central region, the eight strands are associated by I-motif of double A-quartets. Furthermore, eight hydrated-magnesium cations link the four duplexes to support the octaplex formation. Based on these structural features, a proposal that folding of d(GAAA)n, found in the non-coding region of genomes, into an octaplex can induce slippage during replication to facilitate length polymorphism is presented.
- Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, 4259 Nagatsuda, Midori-ku, Yokohama 226-850.
Organizational Affiliation: