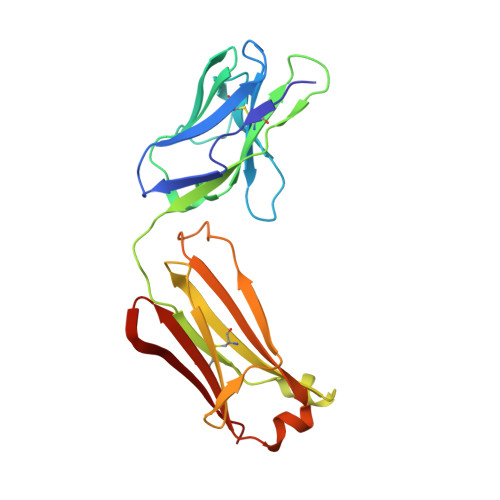
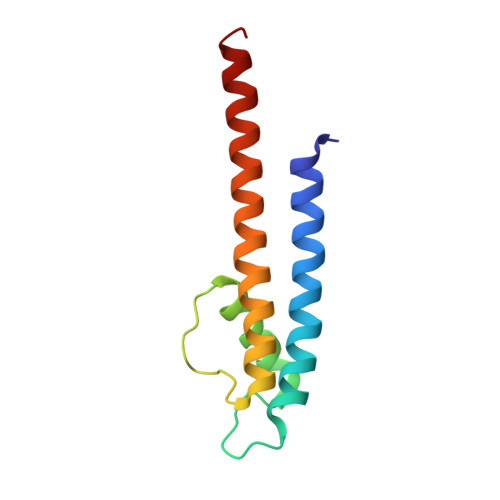
Crystallographic Study of the Tetrabutylammonium Block to the KcsA K(+) Channel
Yohannan, S., Hu, Y., Zhou, Y.(2007) J Mol Biology 366: 806-814
- PubMed: 17196615 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.11.081
- Primary Citation Related Structures:
2DWD, 2DWE, 2HVJ, 2HVK - PubMed Abstract:
K(+) channels play essential roles in regulating membrane excitability of many diverse cell types by selectively conducting K(+) ions through their pores. Many diverse molecules can plug the pore and modulate the K(+) current. Quaternary ammonium (QA) ions are a class of pore blockers that have been used for decades by biophysicists to probe the pore, leading to important insights into the structure-function relation of K(+) channels. However, many key aspects of the QA-blocking mechanisms remain unclear to date, and understanding these questions requires high resolution structural information. Here, we address the question of whether intracellular QA blockade causes conformational changes of the K(+) channel selectivity filter. We have solved the structures of the KcsA K(+) channel in complex with tetrabutylammonium (TBA) and tetrabutylantimony (TBSb) under various ionic conditions. Our results demonstrate that binding of TBA or TBSb causes no significant change in the KcsA structure at high concentrations of permeant ions. We did observe the expected conformational change of the filter at low concentration of K(+), but this change appears to be independent of TBA or TBSb blockade.
- Department of Cellular and Molecular Physiology, Yale University School of Medicine, New Haven, CT 06520, USA.
Organizational Affiliation: