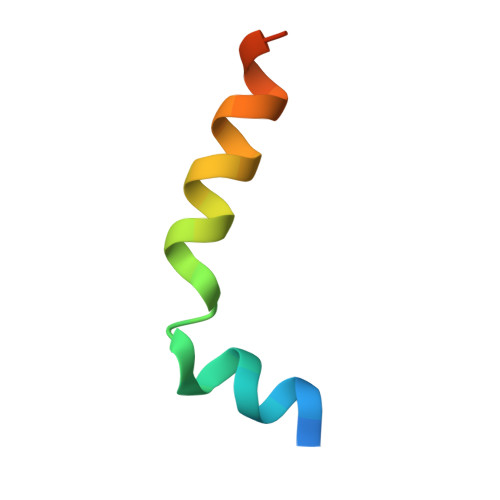
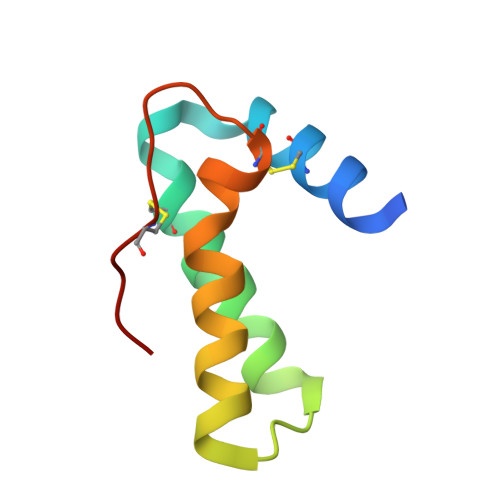
Crystal structure of Mabinlin II: a novel structural type of sweet proteins and the main structural basis for its sweetness.
Li, D.F., Jiang, P., Zhu, D.Y., Hu, Y., Max, M., Wang, D.C.(2008) J Struct Biol 162: 50-62
- PubMed: 18308584 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2007.12.007
- Primary Citation Related Structures:
2DS2 - PubMed Abstract:
The crystal structure of a sweet protein Mabinlin II (Mab II) isolated from the mature seeds of Capparis masaikai Levl. grown in Southern China has been determined at 1.7A resolution by the SIRAS method. The Mab II 3D structure features in an "all alpha" fold mode consisting of A- and B-chains crosslinked by four disulfide bridges, which is distinct from all known sweet protein structures. The Mabinlin II molecule shows an amphiphilic surface, a cationic face (Face A) and a neutral face (Face B). A unique structural motif consisting of B54-B64 was found in Face B, which adopts a special sequence, NL-P-NI-C-NI-P-NI, featuring four [Asn-Leu/Ile] units connected by three conformational-constrained residues, thus is called the [NL/I] tetralet motif. The experiments for testing the possible interactions of separated A-chain and B-chain and the native Mabinlin II to the sweet-taste receptor were performed through the calcium imaging experiments with the HEK293E cells coexpressed hT1R2/T1R3. The result shows that hT1R2/T1R3 responds to both the integrated Mabinlin II and the individual B-chain in the same scale, but not to A-chain. The sweetness evaluation further identified that the separated B-chain can elicit the sweetness alone, but A-chain does not. All data in combination revealed that the sweet protein Mabinlin II can interact with the sweet-taste receptor hT1R2/T1R3 to elicit its sweet taste, and the B-chain with a unique [NL/I] tetralet motif is the essential structural element for the interaction with sweet-taste receptor to elicit the sweetness, while the A-chain may play a role in gaining a long aftertaste for the integrate Mabinlin II. The findings reported in this paper will be advantage for understanding the diversity of sweet proteins and engineering research for development of a unique sweetener for the food and agriculture based on the Mabinlin II structure as a native model.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, People's Republic of China.
Organizational Affiliation: