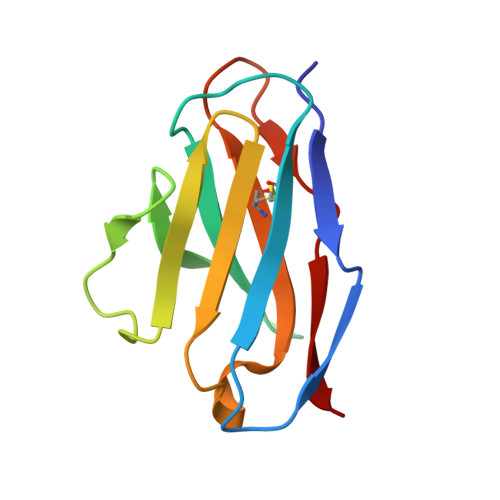
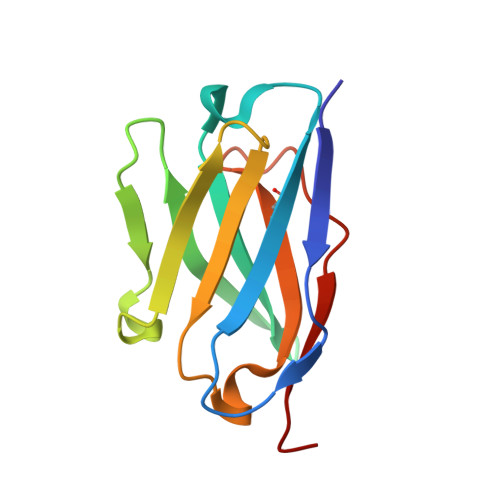
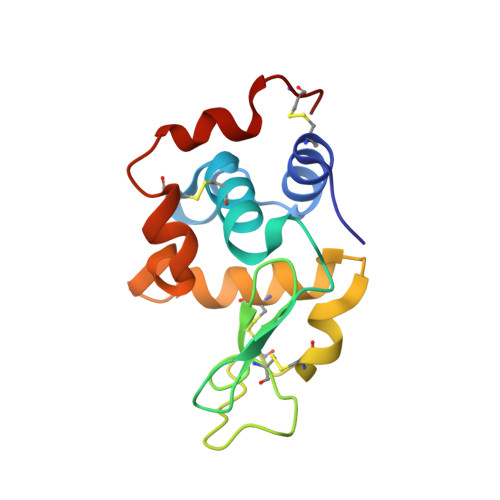
Structural consequences of mutations in interfacial Tyr residues of a protein antigen-antibody complex. The case of HyHEL-10-HEL
Shiroishi, M., Tsumoto, K., Tanaka, Y., Yokota, A., Nakanishi, T., Kondo, H., Kumagai, I.(2007) J Biological Chem 282: 6783-6791
- PubMed: 17166830 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M605197200
- Primary Citation Related Structures:
2DQC, 2DQD, 2DQE, 2DQF, 2DQG, 2DQH, 2DQI, 2DQJ - PubMed Abstract:
Tyrosine is an important amino acid in protein-protein interaction hot spots. In particular, many Tyr residues are located in the antigen-binding sites of antibodies and endow high affinity and high specificity to these antibodies. To investigate the role of interfacial Tyr residues in protein-protein interactions, we performed crystallographic studies and thermodynamic analyses of the interaction between hen egg lysozyme (HEL) and the anti-HEL antibody HyHEL-10 Fv fragment. HyHEL-10 has six Tyr residues in its antigen-binding site, which were systematically mutated to Phe and Ala using site-directed mutagenesis. The crystal structures revealed several critical roles for these Tyr residues in the interaction between HEL and HyHEL-10 as follows: 1) the aromatic ring of Tyr-50 in the light chain (LTyr-50) was important for the correct ternary structure of variable regions of the immunoglobulin light chain and heavy chain and of HEL; 2) deletion of the hydroxyl group of Tyr-50 in the heavy chain (HTyr-50) resulted in structural changes in the antigen-antibody interface; and 3) the side chains of HTyr-33 and HTyr-53 may help induce fitting of the antibody to the antigen. Hot spot Tyr residues may contribute to the high affinity and high specificity of the antigen-antibody interaction through a diverse set of structural and thermodynamic interactions.
- Department of Biomolecular Engineering, Graduate School of Engineering, Tohoku University, Aoba-yama 6-6-11, Sendai 980-8579, Japan.
Organizational Affiliation: