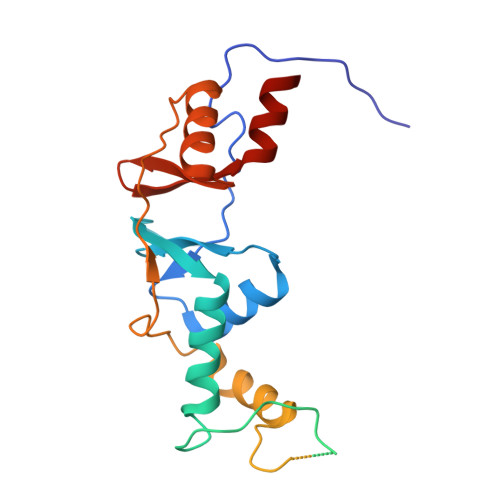
Crystal structure of the cytoplasmic domain of the chloride channel ClC-0.
Meyer, S., Dutzler, R.(2006) Structure 14: 299-307
- PubMed: 16472749 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.10.008
- Primary Citation Related Structures:
2D4Z - PubMed Abstract:
Ion channels are frequently organized in a modular fashion and consist of a membrane-embedded pore domain and a soluble regulatory domain. A similar organization is found for the ClC family of Cl- channels and transporters. Here, we describe the crystal structure of the cytoplasmic domain of ClC-0, the voltage-dependent Cl- channel from T. marmorata. The structure contains a folded core of two tightly interacting cystathionine beta-synthetase (CBS) subdomains. The two subdomains are connected by a 96 residue mobile linker that is disordered in the crystals. As revealed by analytical ultracentrifugation, the domains form dimers, thereby most likely extending the 2-fold symmetry of the transmembrane pore. The structure provides insight into the organization of the cytoplasmic domains within the ClC family and establishes a framework for guiding future investigations on regulatory mechanisms.
- Department of Biochemistry, University of Zürich, Winterthurer Strasse 190, CH-8057 Zürich, Switzerland.
Organizational Affiliation: