Efficient Leukocyte Ig-like Receptor Signaling and Crystal Structure of Disulfide-linked HLA-G Dimer
Shiroishi, M., Kuroki, K., Ose, T., Rasubala, L., Shiratori, I., Arase, H., Tsumoto, K., Kumagai, I., Kohda, D., Maenaka, K.(2006) J Biological Chem 281: 10439-10447
- PubMed: 16455647 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M512305200
- Primary Citation Related Structures:
2D31 - PubMed Abstract:
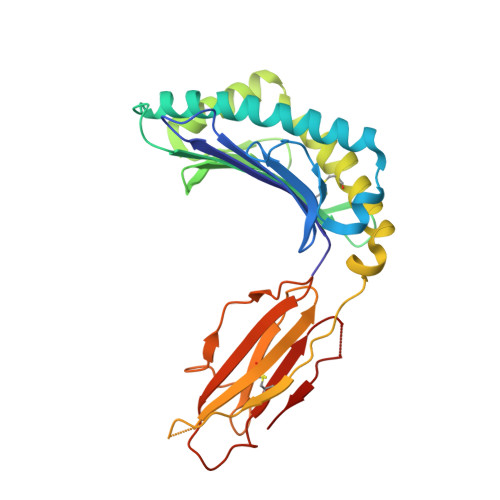
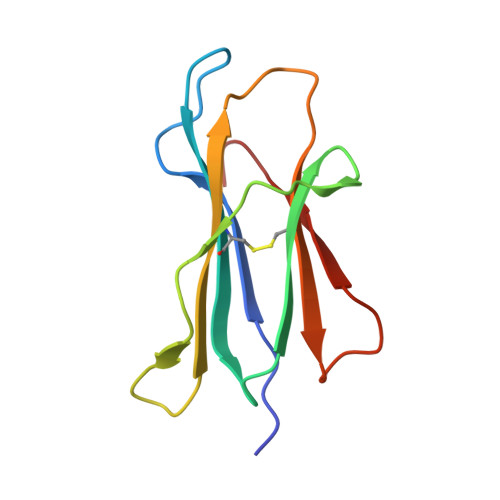
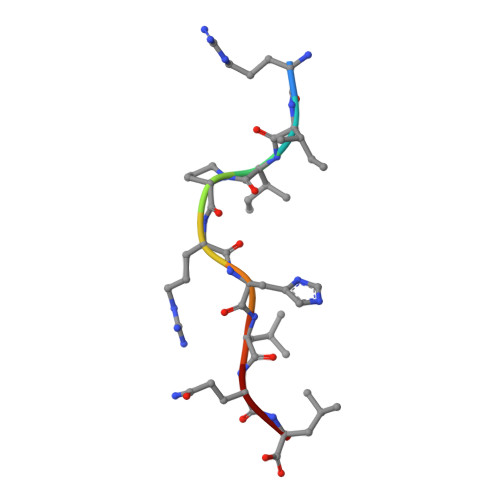
HLA-G is a nonclassical major histocompatibility complex class I (MHCI) molecule, which is expressed in trophoblasts and confers immunological tolerance in the maternal-fetal interface by binding to leukocyte Ig-like receptors (LILRs, also called as LIR/ILT/CD85) and CD8. HLA-G is expressed in disulfide-linked dimer form both in solution and at the cell surface. Interestingly, MHCI dimer formations have been involved in pathogenesis and T cell activation. The structure and receptor binding characteristics of MHCI dimers have never been evaluated. Here we performed binding studies showing that the HLA-G dimer exhibited higher overall affinity to LILRB1/2 than the monomer by significant avidity effects. Furthermore, the cell reporter assay demonstrated that the dimer formation remarkably enhanced the LILRB1-mediated signaling at the cellular level. We further determined the crystal structure of the wild-type dimer of HLA-G with the intermolecular Cys(42)-Cys(42) disulfide bond. This dimer structure showed the oblique configuration to expose two LILR/CD8-binding sites upward from the membrane easily accessible for receptors, providing plausible 1:2 (HLA-G dimer:receptors) complex models. These results indicated that the HLA-G dimer conferred increased avidity in a proper structural orientation to induce efficient LILR signaling, resulting in the dominant immunosuppressive effects. Moreover, structural and functional implications for other MHCI dimers observed in activated T cells and the pathogenic allele, HLA-B27, are discussed.
- Division of Structural Biology, Medical Institute of Bioregulation, Kyushu University, 3-1-1 Maidashi, Higashi-ku, Fukuoka 812-8582, Japan.
Organizational Affiliation: