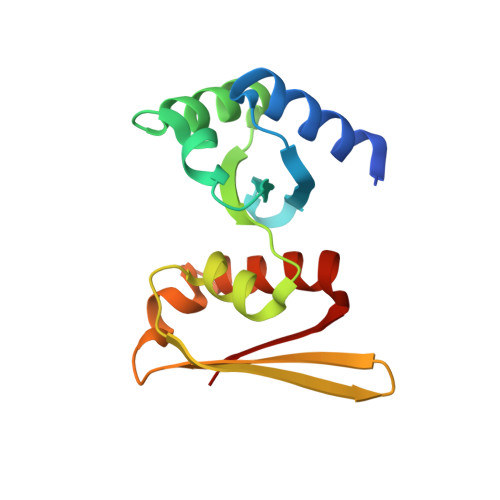
Crystal structure and RNA-binding analysis of the archaeal transcription factor NusA
Shibata, R., Bessho, Y., Shinkai, A., Nishimoto, M., Fusatomi, E., Terada, T., Shirouzu, M., Yokoyama, S.(2007) Biochem Biophys Res Commun 355: 122-128
- PubMed: 17288993 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2007.01.119
- Primary Citation Related Structures:
2CXC - PubMed Abstract:
The transcription factor NusA functions in transcriptional regulation involving termination in bacteria. A NusA homolog consisting of only the two KH domains is widely conserved in archaea, but its function remains unknown. We have found that Aeropyrum pernix NusA strongly binds to a certain CU-rich sequence near a termination signal. Our crystal structure of A. pernix NusA revealed that its spatial arrangement is quite similar to that of the KH domains of bacterial NusA. Thus, we consider archaeal NusA to have retained some functions of bacterial NusA, including the ssRNA-binding ability. Remarkable structural differences between archaeal and bacterial NusA exist at the interface with RNAP, in connection with the different NusA-binding sites around the termination signals. Transcriptional termination in archaea could differ from all of the known bacterial and eukaryal mechanisms, in terms of the combination of a bacterial factor and a eukaryal-type RNAP.
- RIKEN Genomic Sciences Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: