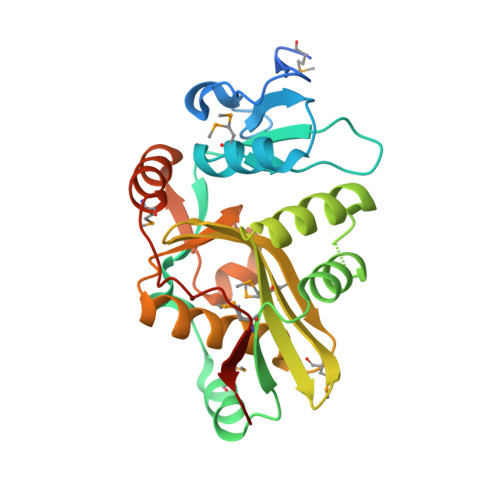
The crystal structure of leucyl/phenylalanyl-tRNA-protein transferase from Escherichia coli
Dong, X., Kato-Murayama, M., Muramatsu, T., Mori, H., Shirouzu, M., Bessho, Y., Yokoyama, S.(2007) Protein Sci 16: 528-534
- PubMed: 17242373 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062616107
- Primary Citation Related Structures:
2CXA - PubMed Abstract:
Leucyl/phenylalanyl-tRNA-protein transferase (L/F-transferase) is an N-end rule pathway enzyme, which catalyzes the transfer of Leu and Phe from aminoacyl-tRNAs to exposed N-terminal Arg or Lys residues of acceptor proteins. Here, we report the 1.6 A resolution crystal structure of L/F-transferase (JW0868) from Escherichia coli, the first three-dimensional structure of an L/F-transferase. The L/F-transferase adopts a monomeric structure consisting of two domains that form a bilobate molecule. The N-terminal domain forms a small lobe with a novel fold. The large C-terminal domain has a highly conserved fold, which is observed in the GCN5-related N-acetyltransferase (GNAT) family. Most of the conserved residues of L/F-transferase reside in the central cavity, which exists at the interface between the N-terminal and C-terminal domains. A comparison of the structures of L/F-transferase and the bacterial peptidoglycan synthase FemX, indicated a structural homology in the C-terminal domain, and a similar domain interface region. Although the peptidyltransferase function is shared between the two proteins, the enzymatic mechanism would differ. The conserved residues in the central cavity of L/F-transferase suggest that this region is important for the enzyme catalysis.
- RIKEN Genomic Sciences Center, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: