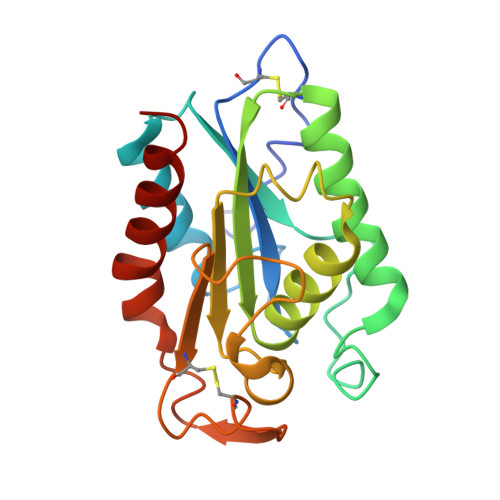
Cutinase, a lipolytic enzyme with a preformed oxyanion hole.
Martinez, C., Nicolas, A., van Tilbeurgh, H., Egloff, M.P., Cudrey, C., Verger, R., Cambillau, C.(1994) Biochemistry 33: 83-89
- PubMed: 8286366 Search on PubMed
- DOI: https://doi.org/10.1021/bi00167a011
- Primary Citation Related Structures:
2CUT - PubMed Abstract:
Cutinases, a group of cutin degrading enzymes with molecular masses of around 22-25 kDa (Kolattukudy, 1984), are also able to efficiently hydrolyse triglycerides (De Geus et al., 1989; Lauwereys et al., 1991), but without exhibiting the interfacial activation phenomenom (Sarda et al., 1958). They belong to a class of proteins with a common structural framework, called the alpha/beta hydrolase fold (Martinez et al., 1992; Ollis et al., 1992). We describe herein the structure of cutinase covalently inhibited by diethyl-p-nitrophenyl phosphate (E600) and refined at 1.9-A resolution. Contrary to what has previously been reported with lipases (Brzozowski et al., 1991; Derewenda et al., 1992; Van Tilbeurgh et al., 1993), no significant structural rearrangement was observed here in cutinase upon the inhibitor binding. Moreover, the structure of the active site machinery, consisting of a catalytic triad (S120, H188, D175) and an oxyanion hole (Q121 and S42), was found to be identical to that of the native enzyme, whereas the oxyanion hole of Rhizomucor lipase (Brzozowski et al., 1991; Derewenda et al., 1992), like that of pancreatic lipase (van Tilbeurgh et al., 1993), is formed only upon enzyme-ligand complex formation. The fact that cutinase does not display interfacial activation cannot therefore only be due to the absence of a lid but might also be attributable to the presence of a preformed oxyanion hole.
- Laboratoire de Cristallisation et Cristallographie des Macromolécules Biologiques, URA 1296-CNRS, Faculté de Médecine Nord, Marseille, France.
Organizational Affiliation: