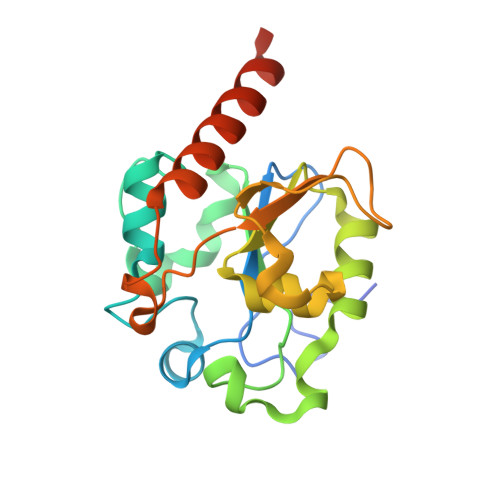
The Crystal Structure of Mismatch Specific Uracil-DNA Glycosylase (Mug) from Deinococcus Radiodurans Reveals a Novel Catalytic Residue and Broad Substrate Specificity
Moe, E., Leiros, I., Smalas, A.O., Mcsweeney, S.(2006) J Biological Chem 281: 569
- PubMed: 16223719 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M508032200
- Primary Citation Related Structures:
2C2P, 2C2Q - PubMed Abstract:
Deinococcus radiodurans is extremely resistant to the effects of ionizing radiation. The source of the radiation resistance is not known, but an expansion of specific protein families related to stress response and damage control has been observed. DNA repair enzymes are among the expanded protein families in D. radiodurans, and genes encoding five different uracil-DNA glycosylases are identified in the genome. Here we report the three-dimensional structure of the mismatch-specific uracil-DNA glycosylase (MUG) from D. radiodurans (drMUG) to a resolution of 1.75 angstroms. Structural analyses suggest that drMUG possesses a novel catalytic residue, Asp-93. Activity measurements show that drMUG has a modified and broadened substrate specificity compared with Escherichia coli MUG. The importance of Asp-93 for activity was confirmed by structural analysis and abolished activity for the mutant drMUGD93A. Two other microorganisms, Bradyrhizobium japonicum and Rhodopseudomonas palustris, possess genes that encode MUGs with the highest sequence identity to drMUG among all of the bacterial MUGs examined. A phylogenetic analysis indicates that these three MUGs form a new MUG/thymidine-DNA glycosylase subfamily, here called the MUG2 family. We suggest that the novel catalytic residue (Asp-93) has evolved to provide drMUG with broad substrate specificity to increase the DNA repair repertoire of D. radiodurans.
- Norwegian Structural Biology Centre, University of Tromsø, N-9037 Tromsø, Norway.
Organizational Affiliation: