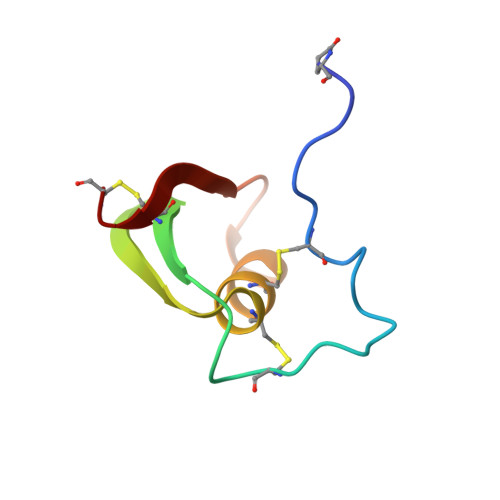
Solution conformation of proteinase inhibitor IIA from bull seminal plasma by 1H nuclear magnetic resonance and distance geometry.
Williamson, M.P., Havel, T.F., Wuthrich, K.(1985) J Mol Biology 182: 295-315
- PubMed: 3839023 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(85)90347-x
- Primary Citation Related Structures:
1BUS, 2BUS - PubMed Abstract:
A determination of the solution conformation of the proteinase inhibitor IIA from bull seminal plasma (BUSI IIA) is described. Two-dimensional nuclear Overhauser enhancement spectroscopy (NOESY) was used to obtain a list of 202 distance constraints between individually assigned hydrogen atoms of the polypeptide chain, to identify the positions of the three disulfide bridges, and to locate the single cis peptide bond. Supplementary geometric constraints were derived from the vicinal spin-spin couplings and the locations of certain hydrogen bonds, as determined by nuclear magnetic resonance (n.m.r.). Using a new distance geometry program (DISGEO) which is capable of computing all-atom structures for proteins the size of BUSI IIA, five conformers were computed from the NOE distance constraints alone, and another five were computed with the supplementary constraints included. Comparison of the different structures computed from the n.m.r. data among themselves and with the crystal structures of two homologous proteins shows that the global features of the conformation of BUSI IIA (i.e. the overall dimensions of the molecule and the threading of the polypeptide chain) were well-defined by the available n.m.r. data. In the Appendix, we describe a preliminary energy refinement of the structure, which showed that the constraints derived from the n.m.r. data are compatible with a low energy spatial structure.