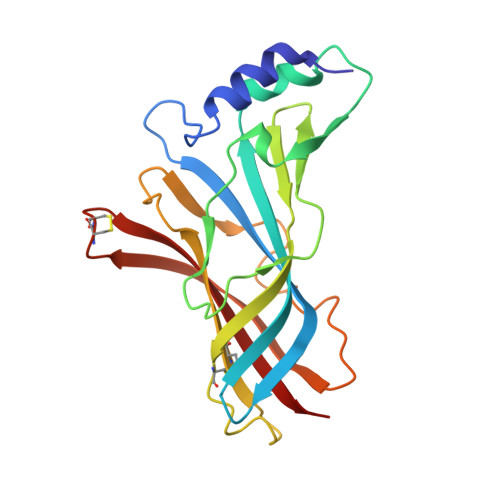
Crystal Structure of Acetylcholine-Binding Protein from Bulinus Truncatus Reveals the Conserved Structural Scaffold and Sites of Variation in Nicotinic Acetylcholine Receptors.
Celie, P.H.N., Klaassen, R.V., Van Rossum-Fikkert, S.E., Van Elk, R., Van Nierop, P., Smit, A.B., Sixma, T.K.(2005) J Biological Chem 280: 26457
- PubMed: 15899893 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M414476200
- Primary Citation Related Structures:
2BJ0 - PubMed Abstract:
The crystal structure of acetylcholine-binding protein (AChBP) from the mollusk Lymnaea stagnalis is the established model for the ligand binding domains of the ligand-gated ion channel family, which includes nicotinic acetylcholine, 5-hydroxytryptamine (5-HT3), gamma-aminobutyric acid (GABA), types A and C, and glycine receptors. Here we present the crystal structure of a remote homolog, AChBP from Bulinus truncatus, which reveals both the conserved structural scaffold and the sites of variation in this receptor family. These include rigid body movements of loops that are close to the transmembrane interface in the receptors and changes in the intermonomer contacts, which alter the pentamer stability drastically. Structural, pharmacological and mutational analysis of both AChBPs shows how 3 amino acid changes in the binding site contribute to a 5-10-fold difference in affinity for nicotinic ligands. Comparison of these structures will be valuable for improving structure-function studies of ligand-gated ion channel receptors, including signal transduction, homology modeling, and drug design.
- Division of Molecular Carcinogenesis, Netherlands Cancer Institute, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands.
Organizational Affiliation: