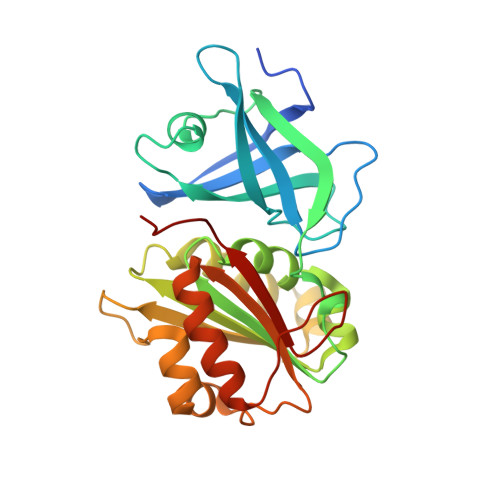
The ferredoxin-NADP(H) reductase from Rhodobacter capsulatus: molecular structure and catalytic mechanism.
Nogues, I., Perez-Dorado, I., Frago, S., Bittel, C., Mayhew, S.G., Gomez-Moreno, C., Hermoso, J.A., Medina, M., Cortez, N., Carrillo, N.(2005) Biochemistry 44: 11730-11740
- PubMed: 16128574 Search on PubMed
- DOI: https://doi.org/10.1021/bi0508183
- Primary Citation Related Structures:
2BGI, 2BGJ - PubMed Abstract:
The photosynthetic bacterium Rhodobacter capsulatus contains a ferredoxin (flavodoxin)-NADP(H) oxidoreductase (FPR) that catalyzes electron transfer between NADP(H) and ferredoxin or flavodoxin. The structure of the enzyme, determined by X-ray crystallography, contains two domains harboring the FAD and NADP(H) binding sites, as is typical of the FPR structural family. The FAD molecule is in a hairpin conformation in which stacking interactions can be established between the dimethylisoalloxazine and adenine moieties. The midpoint redox potentials of the various transitions undergone by R. capsulatus FPR were similar to those reported for their counterparts involved in oxygenic photosynthesis, but its catalytic activity is orders of magnitude lower (1-2 s(-)(1) versus 200-500 s(-)(1)) as is true for most of its prokaryotic homologues. To identify the mechanistic basis for the slow turnover in the bacterial enzymes, we dissected the R. capsulatus FPR reaction into hydride transfer and electron transfer steps, and determined their rates using stopped-flow methods. Hydride exchange between the enzyme and NADP(H) occurred at 30-150 s(-)(1), indicating that this half-reaction does not limit FPR activity. In contrast, electron transfer to flavodoxin proceeds at 2.7 s(-)(1), in the range of steady-state catalysis. Flavodoxin semiquinone was a better electron acceptor for FPR than oxidized flavodoxin under both single turnover and steady-state conditions. The results indicate that one-electron reduction of oxidized flavodoxin limits the enzyme activity in vitro, and support the notion that flavodoxin oscillates between the semiquinone and fully reduced states when FPR operates in vivo.
- Departamento de Bioquímica y Biología Molecular y Celular, Facultad de Ciencias, and Institute of Biocomputation and Physics of Complex Systems (BIFI), Universidad de Zaragoza, Zaragoza, Spain.
Organizational Affiliation: