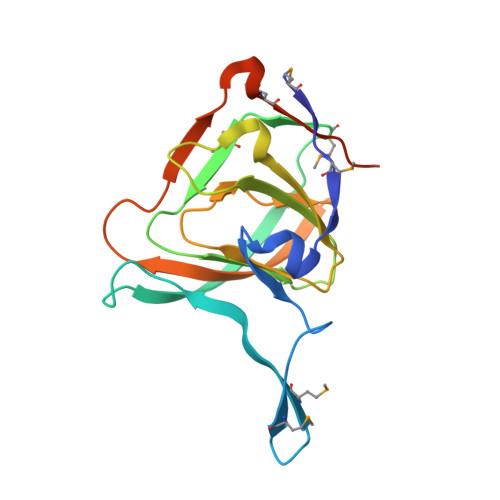
Crystal Structure of the Putative Ureidoglycolate hydrolase PP4288 from Pseudomonas putida, Northeast Structural Genomics Target PpR49
Forouhar, F., Abashidze, M., Jayaraman, S., Ho, C.K., Conover, K., Acton, T.B., Montelione, G.T., Hunt, J.F., Tong, L.To be published.