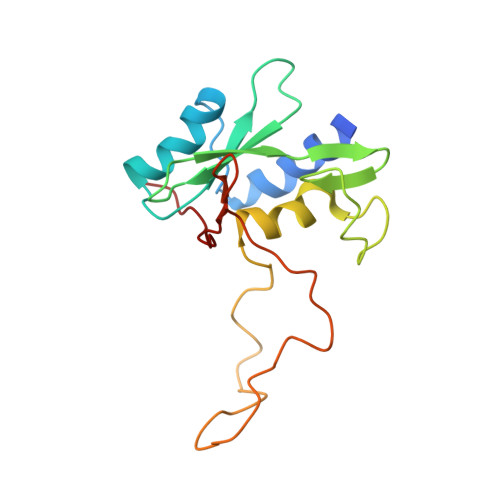
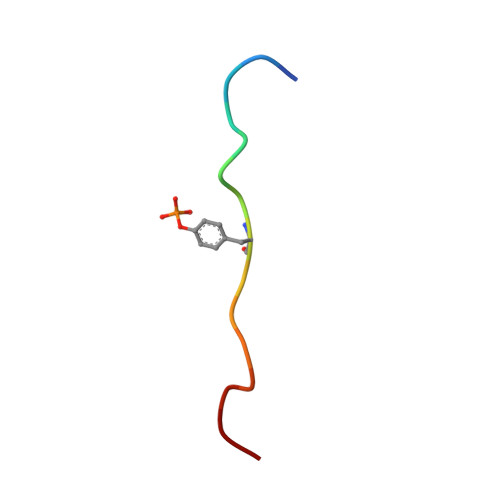
The Structure of SOCS3 Reveals the Basis of the Extended SH2 Domain Function and Identifies an Unstructured Insertion That Regulates Stability
Babon, J.J., McManus, E.J., Yao, S., DeSouza, D.P., Mielke, L.A., Sprigg, N.S., Willson, T.A., Hilton, D.J., Nicola, N.A., Baca, M., Nicholson, S.E., Norton, R.S.(2006) Mol Cell 22: 205-216
- PubMed: 16630890 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2006.03.024
- Primary Citation Related Structures:
2BBU - PubMed Abstract:
SOCS3 is essential for regulating the extent, duration, and specificity of cellular responses to cytokines such as G-CSF and IL-6. Here we describe the solution structure of SOCS3, the first structure determined for any SOCS protein, in complex with a phosphotyrosine-containing peptide from the IL-6 receptor signaling subunit gp130. The structure of the complex shows that seven peptide residues form a predominantly hydrophobic binding motif. Regions outside the SOCS3 SH2 domain are important for ligand binding, in particular, a single 15 residue alpha helix immediately N-terminal to the SH2 domain makes direct contacts with the phosphotyrosine binding loop and, in part, determines its geometry. The SH2 domain itself is remarkable in that it contains a 35 residue unstructured PEST motif insertion that is not required for STAT inhibition. The PEST motif increases SOCS3 turnover and affects its degradation pathway, implying that it has an important regulatory role inside the cell.
- The Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Parkville, 3050, Victoria, Australia. babon@wehi.edu.au
Organizational Affiliation: