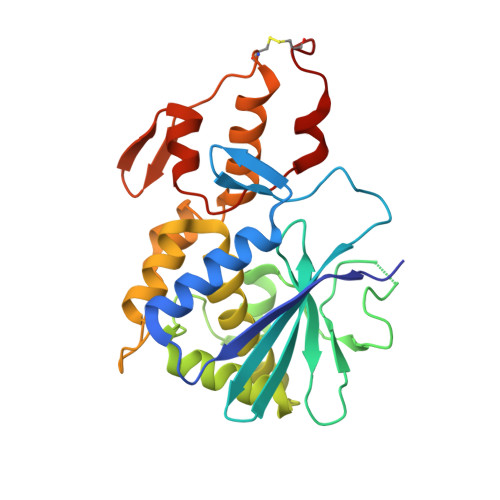
Isolation, characterization, sequencing and crystal structure of charybdin, a type 1 ribosome-inactivating protein from Charybdis maritima agg.
Touloupakis, E., Gessmann, R., Kavelaki, K., Christofakis, E., Petratos, K., Ghanotakis, D.F.(2006) FEBS J 273: 2684-2692
- PubMed: 16817896 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05287.x
- Primary Citation Related Structures:
2B7U - PubMed Abstract:
A novel, type 1 ribosome-inactivating protein designated charybdin was isolated from bulbs of Charybdis maritima agg. The protein, consisting of a single polypeptide chain with a molecular mass of 29 kDa, inhibited translation in rabbit reticulocytes with an IC50 of 27.2 nm. Plant genomic DNA extracted from the bulb was amplified by PCR between primers based on the N-terminal and C-terminal sequence of the protein from dissolved crystals. The complete mature protein sequence was derived by partial DNA sequencing and terminal protein sequencing, and was confirmed by high-resolution crystal structure analysis. The protein contains Val at position 79 instead of the conserved Tyr residue of the ribosome-inactivating proteins known to date. To our knowledge, this is the first observation of a natural substitution of a catalytic residue at the active site of a natural ribosome-inactivating protein. This substitution in the active site may be responsible for the relatively low in vitro translation inhibitory effect compared with other ribosome-inactivating proteins. Single crystals were grown in the cold room from PEG6000 solutions. Diffraction data collected to 1.6 A resolution were used to determine the protein structure by the molecular replacement method. The fold of the protein comprises two structural domains: an alpha + beta N-terminal domain (residues 4-190) and a mainly alpha-helical C-terminal domain (residues 191-257). The active site is located in the interface between the two domains and comprises residues Val79, Tyr117, Glu167 and Arg170.
- Department of Chemistry, University of Crete, Greece.
Organizational Affiliation: