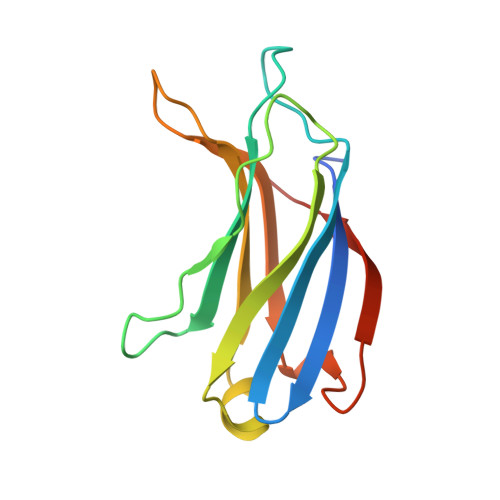
Crystal structure of the C2 domain of class II phosphatidylinositide 3-kinase C2alpha.
Liu, L., Song, X., He, D., Komma, C., Kita, A., Virbasius, J.V., Huang, G., Bellamy, H.D., Miki, K., Czech, M.P., Zhou, G.W.(2006) J Biological Chem 281: 4254-4260
- PubMed: 16338929 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M510791200
- Primary Citation Related Structures:
2B3R - PubMed Abstract:
Phosphatidylinositide (PtdIns) 3-kinase catalyzes the addition of a phosphate group to the 3'-position of phosphatidyl inositol. Accumulated evidence shows that PtdIns 3-kinase can provide a critical signal for cell proliferation, cell survival, membrane trafficking, glucose transport, and membrane ruffling. Mammalian PtdIns 3-kinases are divided into three classes based on structure and substrate specificity. A unique characteristic of class II PtdIns 3-kinases is the presence of both a phox homolog domain and a C2 domain at the C terminus. The biological function of the C2 domain of the class II PtdIns 3-kinases remains to be determined. We have determined the crystal structure of the mCPK-C2 domain, which is the first three-dimensional structural model of a C2 domain of class II PtdIns 3-kinases. Structural studies reveal that the mCPK-C2 domain has a typical anti-parallel beta-sandwich fold. Scrutiny of the surface of this C2 domain has identified three small, shallow sulfate-binding sites. On the basis of the structural features of these sulfate-binding sites, we have studied the lipid binding properties of the mCPK-C2 domain by site-directed mutagenesis. Our results show that this C2 domain binds specifically to PtdIns(3,4)P(2) and PtdIns(4,5)P(2) and that three lysine residues at SBS I site, Lys-1420, Lys-1432, and Lys-1434, are responsible for the phospholipid binding affinity.
- Department of Biological Sciences, Louisiana State University, Baton Rouge, 70803, USA.
Organizational Affiliation: