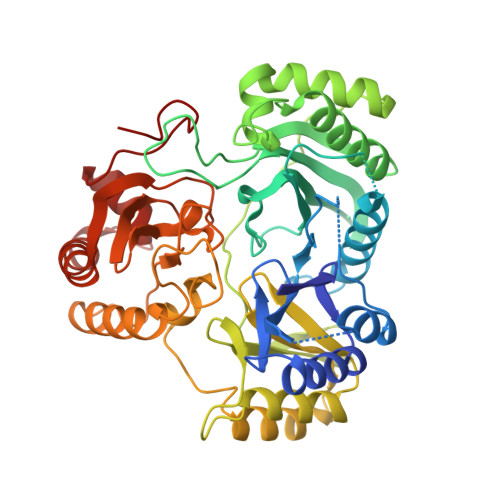
Structures of the siroheme- and Fe4S4-containing active center of sulfite reductase in different states of oxidation: heme activation via reduction-gated exogenous ligand exchange.
Crane, B.R., Siegel, L.M., Getzoff, E.D.(1997) Biochemistry 36: 12101-12119
- PubMed: 9315848 Search on PubMed
- DOI: https://doi.org/10.1021/bi971065q
- Primary Citation Related Structures:
2AOP, 3AOP, 4AOP, 5AOP - PubMed Abstract:
The active center of the Escherichia coli sulfite reductase hemoprotein (SiRHP) is exquisitely designed to catalyze the six-electron reductions of sulfite to sulfide and nitrite to ammonia. Refined high-resolution crystallographic structures of oxidized, two-electron reduced, and intermediately reduced states of SiRHP, monitored by single-crystal electron paramagnetic resonance (EPR) spectroscopy, reveal that a bridging cysteine thiolate supplied by the protein always covalently links the siroheme (iron isobacteriochlorin) to the Fe4S4 cluster, facilitating their ability to transfer electrons to substrate. The reduction potential and reactivity of the cluster are tuned by association with the siroheme, accessibility to solvent, and hydrogen bonds supplied by the protein loops containing the four cluster-ligating cysteines. The distorted conformation of the siroheme recognized by the protein potentially destabilizes the electronic conjugation of the isobacteriochlorin ring and produces axial configurations for some propionate side chains that promote interactions with exogenous ligands and active-site residues. An extensive hydrogen-bond network of positively charged side chains, ordered water molecules, and siroheme carboxylates coordinates, polarizes, and influences the protonation state of anionic ligands. In the oxidized (siroheme Fe3+, Fe4S42+) SiRHP crystal structure, the high density of positive charges in the binding pocket is stabilized by the siroheme's sixth axial ligand-an exogenous phosphate anion. Binding assays with H32PO42- demonstrate that oxidized SiRHP binds phosphate in solution with a dissociation constant of 14 microM at pH 7.7, suggesting that phosphate anions play an important role in stabilizing and sequestering the active-site of the oxidized enzyme in vivo. Reduction of the cofactors couples changes in siroheme iron coordination geometry to changes in active-site protein conformation, leading to phosphate release both in the crystal and in solution. An intermediately reduced enzyme, where the siroheme is mainly ferrous (+2) and the cluster cubane is mainly oxidized (+2), appears to have the lowest affinity for phosphate in the crystal. Reduction-gated release of phosphate from the substrate-binding site may explain the 10(5)-fold increase in rates of ligand association that accompany reduction of SiRHP.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, California 92037, USA.
Organizational Affiliation: