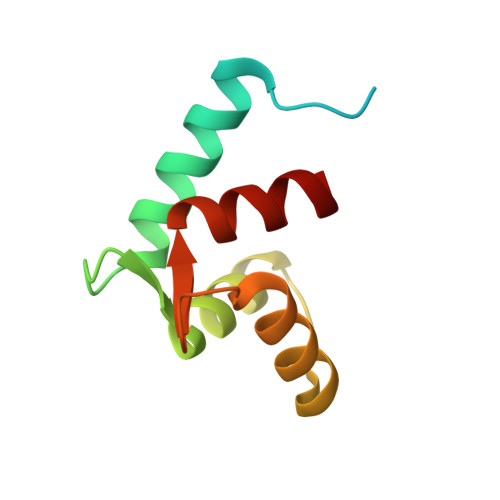
Structure of the N-terminal calcium sensor domain of centrin reveals the biochemical basis for domain-specific function.
Sheehan, J.H., Bunick, C.G., Hu, H., Fagan, P.A., Meyn, S.M., Chazin, W.J.(2006) J Biological Chem 281: 2876-2881
- PubMed: 16317001 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M509886200
- Primary Citation Related Structures:
2AMI - PubMed Abstract:
Centrin is an essential component of microtubule-organizing centers in organisms ranging from algae and yeast to humans. It is an EF-hand calcium-binding protein with homology to calmodulin but distinct calcium binding properties. In a previously proposed model, the C-terminal domain of centrin serves as a constitutive anchor to target proteins, and the N-terminal domain serves as the sensor of calcium signals. The three-dimensional structure of the N-terminal domain of Chlamydomonas rheinhardtii centrin has been determined in the presence of calcium by solution NMR spectroscopy. The domain is found to occupy an open conformation typical of EF-hand calcium sensors. Comparison of the N- and C-terminal domains of centrin reveals a structural and biochemical basis for the domain specificity of interactions with its cellular targets and the distinct nature of centrin relative to other EF-hand proteins. An NMR titration of the centrin N-terminal domain with a fragment of the known centrin target Sfi1 reveals binding of the peptide to a discrete site on the protein, which supports the proposal that the N-terminal domain serves as a calcium sensor in centrin.
- Department of Biochemistry, Vanderbilt University, Nashville, Tennessee 37232-8725, USA.
Organizational Affiliation: