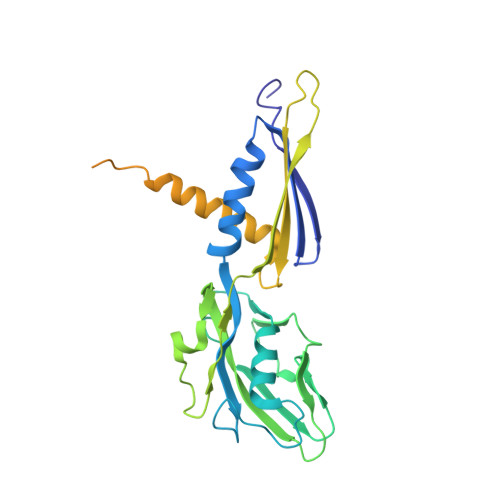
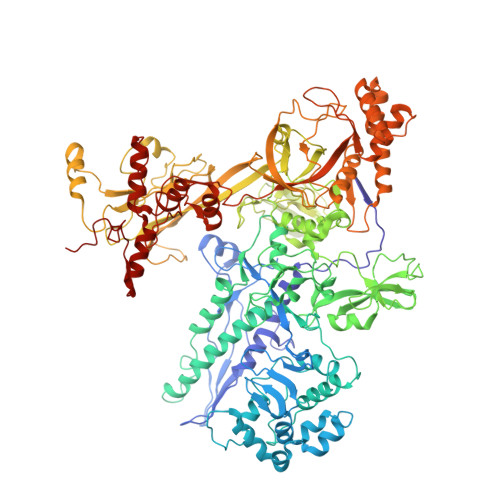
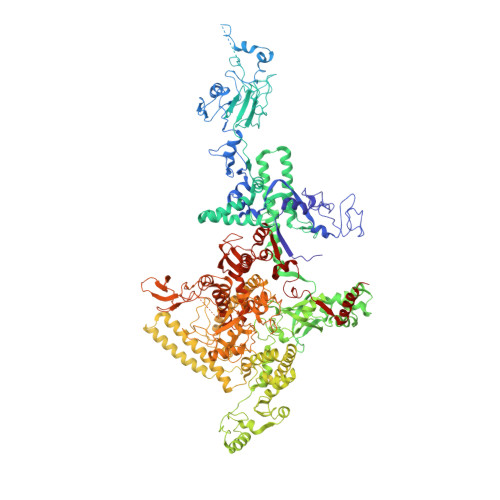
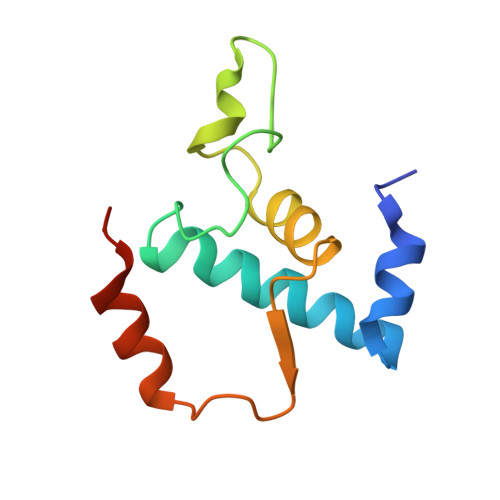
Structural basis of transcription inhibition by antibiotic streptolydigin.
Temiakov, D., Zenkin, N., Vassylyeva, M.N., Perederina, A., Tahirov, T.H., Kashkina, E., Savkina, M., Zorov, S., Nikiforov, V., Igarashi, N., Matsugaki, N., Wakatsuki, S., Severinov, K., Vassylyev, D.G.(2005) Mol Cell 19: 655-666
- PubMed: 16167380 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2005.07.020
- Primary Citation Related Structures:
2A6H - PubMed Abstract:
Streptolydigin (Stl) is a potent inhibitor of bacterial RNA polymerases (RNAPs). The 2.4 A resolution structure of the Thermus thermophilus RNAP-Stl complex showed that, in full agreement with the available genetic data, the inhibitor binding site is located 20 A away from the RNAP active site and encompasses the bridge helix and the trigger loop, two elements that are considered to be crucial for RNAP catalytic center function. Structure-based biochemical experiments revealed additional determinants of Stl binding and demonstrated that Stl does not affect NTP substrate binding, DNA translocation, and phosphodiester bond formation. The RNAP-Stl complex structure, its comparison with the closely related substrate bound eukaryotic transcription elongation complexes, and biochemical analysis suggest an inhibitory mechanism in which Stl stabilizes catalytically inactive (preinsertion) substrate bound transcription intermediate, thereby blocking structural isomerization of RNAP to an active configuration. The results provide a basis for a design of new antibiotics utilizing the Stl-like mechanism.
- Department of Cell Biology, School of Osteopathic Medicine, University of Medicine and Dentistry of New Jersey, Stratford, New Jersey 08084, USA.
Organizational Affiliation: