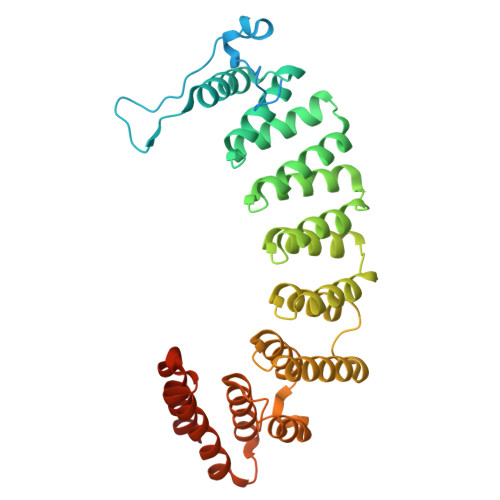
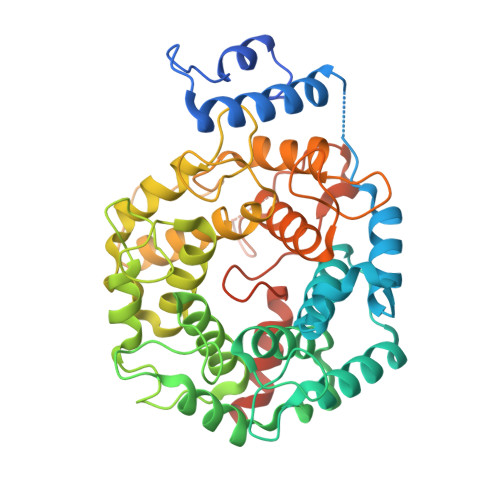
KRAS Can Bind to FTase Despite Disruption of the CAAX Binding Site.
Carion, M., Cuesta, R., Kowalczyk, D., Smets, W., Soons, E., Klaassen, H., Vanderhoydonck, B., Marchand, A., Versele, M., Chaltin, P., Dedecker, P., Park, H., Ismail, S.(2026) Biochemistry
- PubMed: 41680083
- DOI: https://doi.org/10.1021/acs.biochem.5c00732
- Primary Citation of Related Structures:
28JN - PubMed Abstract:
Protein prenylation is a post-translational modification promoting membrane association where isoprenoid lipids attach to C-terminal cysteines of eukaryotic proteins such as Ras and Rho GTPases, nucleus lamins, and G-protein subunits. Three enzymes catalyze this process: farnesyltransferase (FTase) and geranylgeranyltransferase type I and II (GGTase I and RabGGTase). FTase and GGTase-I recognize C-terminal CaaX motifs, of which the terminal amino acid confers specificity. Due to its involvement in oncogenic Ras activation, FTase has become a major anticancer target for drug development. Although first-generation FTase inhibitors failed in clinical trials in many cancers due to compensatory geranylgeranylation of KRAS and NRAS, they remain effective against HRAS-driven tumors and other pathologies, such as Hutchinson-Gilford progeria syndrome. The FTase inhibitor A-176120 was reported to compete with farnesyl and not KRAS. However, our crystallographic and biochemical analyses reveal that A-176120 sterically interferes with the engagement of the KRAS CAAX motif, reducing, but not abolishing, its binding to FTase.
- Department of Chemistry, Biochemistry, Molecular and Structural Biology Division, KU Leuven, 3001 Heverlee, Belgium.
Organizational Affiliation: