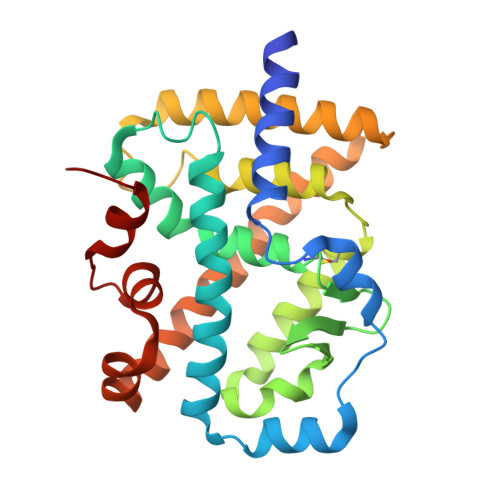
Structure of a chimeric ROR alpha ligand-binding domain in fusion with a RIP-140 coactivator peptide.
Li, X., Zhang, G., Tan, L., Chen, L., Im, Y.J.(2026) Biochem Biophys Res Commun 812: 153622-153622
- PubMed: 41844017 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2026.153622
- Primary Citation Related Structures:
23VY - PubMed Abstract:
The retinoic acid-related orphan receptor α (RORα) is a potential drug target for cancer, inflammation, and metabolic diseases. Structure-guided ligand optimization can facilitate the development of selective RORα modulators. However, structural studies of the RORα ligand-binding domain (LBD) have been limited due to difficulties in purifying recombinant protein suitable for crystallographic analysis. Here, we engineered a chimeric RORα LBD C-terminally fused to the LXXLL motif of the coactivator RIP-140. This fusion improved the stability and solubility of the RORα LBD, enabling high-yield purification using the E. coli expression system. We determined the crystal structure of the chimeric RORα LBD in complex with cholesterol at 2.7 Å resolution. The cholesterol-bound RORα LBD adopted an active conformation of helix 12 that accommodates coactivator binding. The coactivator peptide appears to stabilize the RORα LBD by shielding the hydrophobic surface of the AF-2 region. Although monomeric in solution, the RORα LBD-LXXLL fusion formed a dimer in the crystal lattice through extensive interactions involving the LXXLL motifs and α3 helices, which may represent a physiological homodimer. Fusion of a coactivator motif to the LBD is expected to facilitate structural studies of RORα and may be broadly applicable to other nuclear receptor LBDs for generating stable recombinant proteins.
- College of Pharmacy, Chonnam National University, Gwangju, 61186, Republic of Korea.
Organizational Affiliation: