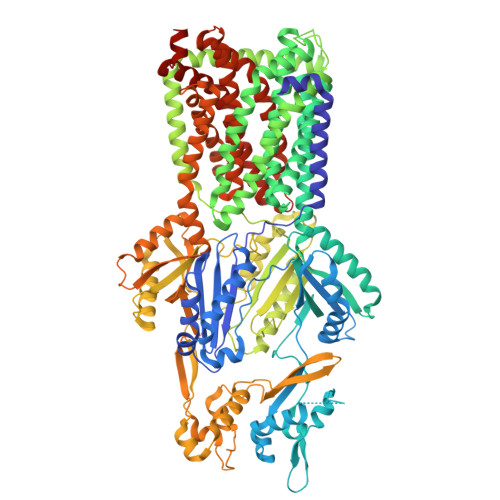
Cryo-EM structures of a MexB-MexY chimeric efflux pump reveal that large open clefts are intrinsic to the MexY porter domain.
Wang, J., Tsutsumi, K., Hirose, M., Nakashima, R., Kato, T., Nishino, K., Nakagawa, A., Yamashita, E.(2026) Acta Crystallogr F Struct Biol Commun 82: 83-93
- PubMed: 41744473 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X26001202
- Primary Citation Related Structures:
22XK, 22XM - PubMed Abstract:
RND-type multidrug-efflux pumps are major contributors to multidrug resistance in Gram-negative bacteria, with MexY from Pseudomonas aeruginosa playing a central role in aminoglycoside resistance. Unlike other RND transporters, MexY exhibits unusually large open clefts in the binding and extrusion states. To determine whether this feature is intrinsic to its drug-recognition porter domain, we created a chimeric protein, MexBYB, by replacing the funnel-like and transmembrane domains of MexY with those of the homologous transporter MexB, and determined its structures by cryoEM under apo and kanamycin-supplemented conditions. Under both conditions, MexBYB was reported to adopt symmetric-like and asymmetric conformations. Structural comparisons reveal that the unusually large open clefts are retained in MexBYB, indicating that this feature is intrinsic to the MexY porter domain.
- Laboratory for Supramolecular Crystallography, Institute for Protein Research, The University of Osaka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: