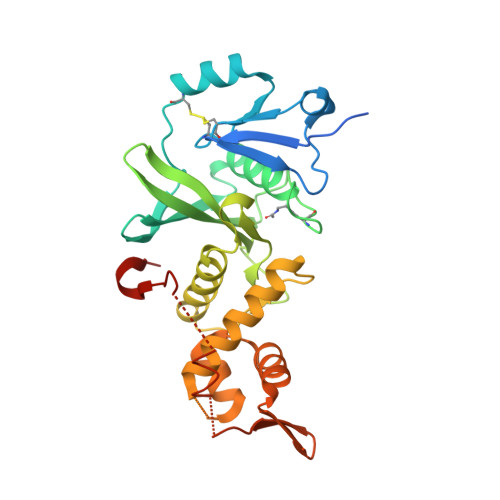
Crystal Structure of MYST histone acetyltransferase KAT6A in complex with Compound 20
NarasimhaRao, K., Vijayshankar, N., SumalathaRani, T., Kalishankar, B., Chandregowda, V., Chandrasekar, A., Susanta, S., Raymond, A.N., David, C.M.To be published.