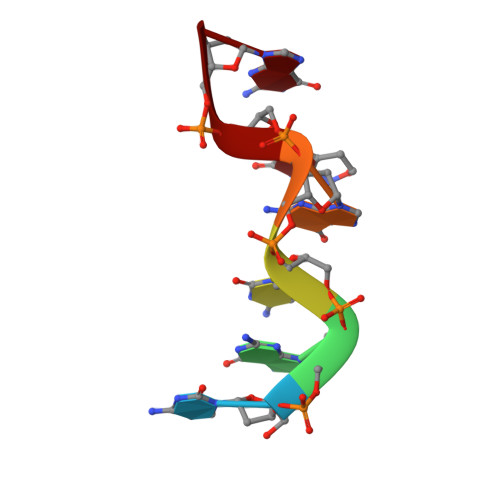
Direct observation of two base-pairing modes of a cytosine-thymine analogue with guanine in a DNA Z-form duplex: significance for base analogue mutagenesis.
Moore, M.H., Van Meervelt, L., Salisbury, S.A., Lin, P.K., Brown, D.M.(1995) J Mol Biology 251: 665-673
- PubMed: 7666418 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0463
- Primary Citation Related Structures:
223D - PubMed Abstract:
The pyrimidine nucleobase analogue 6H,8H-3,4-dihydropyrimido[4,5-c]- [1,2]oxazin-7-one (P) is a mimic both of cytosine and thymine, since it can form stable hydrogen-bonded base-pairs with either guanine or adenine. To investigate the geometric properties of pairing with guanine in a DNA double helix, the structure of d(CGCGPG)2 has been determined by single crystal X-ray analysis. The oligonucleotide crystallised as a left-handed Z-DNA duplex in the orthorhombic space group P2(1)2(1)2(1) with cell dimensions a = 18.23 A, b = 30.63 A, c = 43.78 A. Refinement using NUCLSQ with 51 water molecules included in the final model converged at R = 0.179 (Rw = 0.159) for 2798 reflections (F > 2 sigma (F)) in the range 8 A to 1.7 A. Remarkably, the two P.G pairs in the hexamer duplex are different: Watson-Crick and wobble types separately illustrate both cytosine-like and thymine-like behaviour. The result suggests that mutagenesis experiments involving P and other analogues which display pronounced base-pairing ambivalence can be used to examine the structural basis of substrate discrimination by polymerases that is essential to accurate genetic replication.
- Department of Chemistry, York University, Heslington, UK.
Organizational Affiliation: