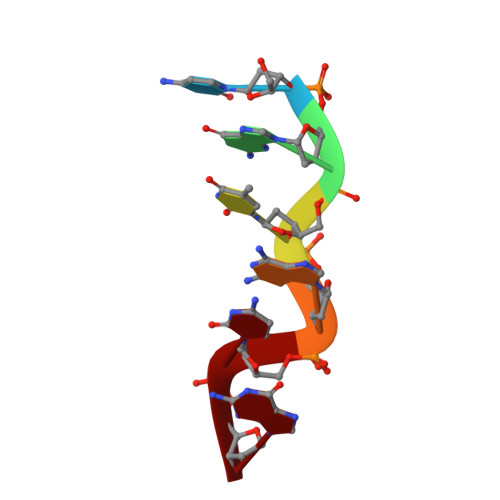
Crystal and molecular structure of a new Z-DNA crystal form: d[CGT(2-NH2-A)CG] and its platinated derivative.
Parkinson, G.N., Arvanitis, G.M., Lessinger, L., Ginell, S.L., Jones, R., Gaffney, B., Berman, H.M.(1995) Biochemistry 34: 15487-15495
- PubMed: 7492550 Search on PubMed
- DOI: https://doi.org/10.1021/bi00047a014
- Primary Citation Related Structures:
210D, 211D - PubMed Abstract:
The three-dimensional structure of d[CGTA'CG], where A' = [2-NH2-A], was determined to atomic (1.35 A) resolution by single isomorphous replacement. The d[CGTA'CG] hexamer crystallizes in space group P3221, and is not isomorphous with other DNA hexanucleotides. Despite completely different crystal packing, the essential characteristics of the Z-DNA conformation are maintained. The structure was determined by single isomorphous replacement using a triammine platinum fragment. Thus, this study also demonstrates, for the first time, the feasibility of the use of this reagent for the direct phasing of DNA crystal structures.
- Department of Chemistry, Rutgers University, Piscataway, New Jersey 08855-0939, USA.
Organizational Affiliation: