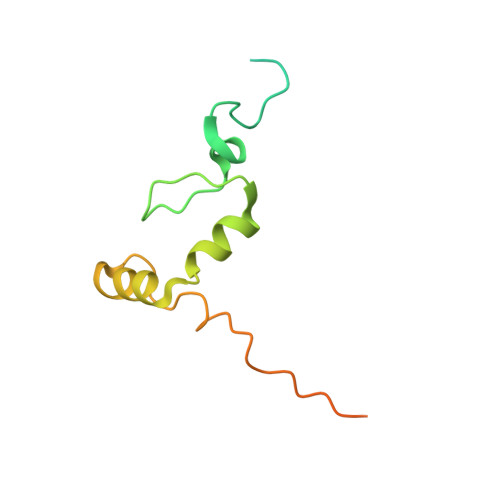
The solution structure of ZNF593 from Homo sapiens reveals a zinc finger in a predominantly unstructured protein.
Hayes, P.L., Lytle, B.L., Volkman, B.F., Peterson, F.C.(2008) Protein Sci 17: 571-576
- PubMed: 18287285 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.073290408
- Primary Citation Related Structures:
1ZR9 - PubMed Abstract:
Here, we report the solution structure of ZNF593, a protein identified in a functional study as a negative modulator of the DNA-binding activity of the Oct-2 transcription factor. ZNF593 contains a classic C(2)H(2) zinc finger domain flanked by about 40 disordered residues on each terminus. Although the protein contains a high degree of intrinsic disorder, the structure of the zinc finger domain was resolved by NMR spectroscopy without a need for N- or C-terminal truncations. The tertiary structure of the zinc finger domain is composed of a beta-hairpin that positions the cysteine side chains for zinc coordination, followed by an atypical kinked alpha-helix containing the two histidine side chain ligands. The structural topology of ZNF593 is similar to a fragment of the double-stranded RNA-binding protein Zfa and the C-terminal zinc finger of MBP-1, a human enhancer binding protein. The structure presented here will provide a guide for future functional studies of how ZNF593 negatively modulates the DNA-binding activity of Oct-2, a POU domain-containing transcription factor. Our work illustrates the unique capacity of NMR spectroscopy for structural analysis of folded domains in a predominantly disordered protein.
- Department of Biochemistry and Center for Eukaryotic Structural Genomics, Medical College of Wisconsin, Milwaukee, Wisconsin 53226, USA.
Organizational Affiliation: