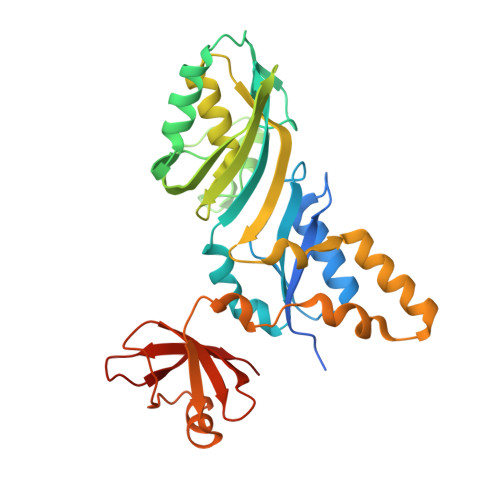
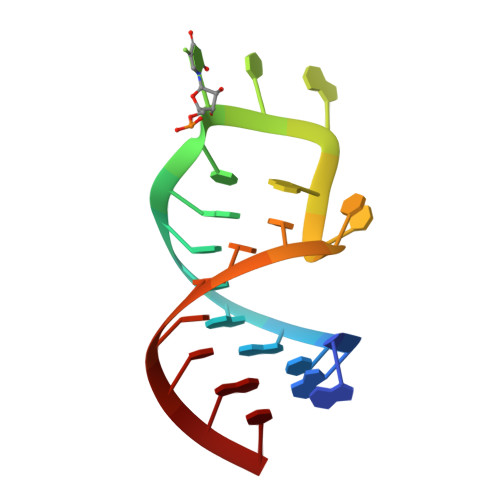
Precursor complex structure of pseudouridine synthase TruB suggests coupling of active site perturbations to an RNA-sequestering peripheral protein domain
Hoang, C., Hamilton, C.S., Mueller, E.G., Ferre-D'Amare, A.R.(2005) Protein Sci 14: 2201-2206
- PubMed: 15987897 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051493605
- Primary Citation Related Structures:
1ZL3 - PubMed Abstract:
The pseudouridine synthase TruB is responsible for the universally conserved post-transcriptional modification of residue 55 of elongator tRNAs. In addition to the active site, the "thumb", a peripheral domain unique to the TruB family of enzymes, makes extensive interactions with the substrate. To coordinate RNA binding and release with catalysis, the thumb may be able to sense progress of the reaction in the active site. To establish whether there is a structural correlate of communication between the active site and the RNA-sequestering thumb, we have solved the structure of a catalytically inactive point mutant of TruB in complex with a substrate RNA, and compared it to the previously determined structure of an active TruB bound to a reaction product. Superposition of the two structures shows that they are extremely similar, except in the active site and, intriguingly, in the relative position of the thumb. Because the two structures were solved using isomorphous crystals, and because the thumb is very well ordered in both structures, the displacement of the thumb we observe likely reflects preferential propagation of active site perturbations to this RNA-binding domain. One of the interactions between the active site and the thumb involves an active site residue whose hydrogen-bonding status changes during the reaction. This may allow the peripheral RNA-binding domain to monitor progress of the pseudouridylation reaction.
- Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, WA 98109-1024, USA.
Organizational Affiliation: