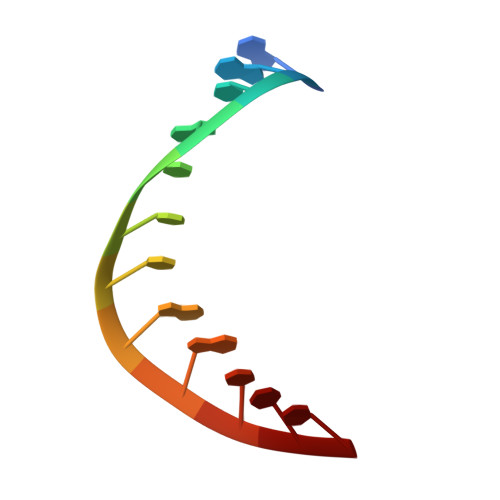
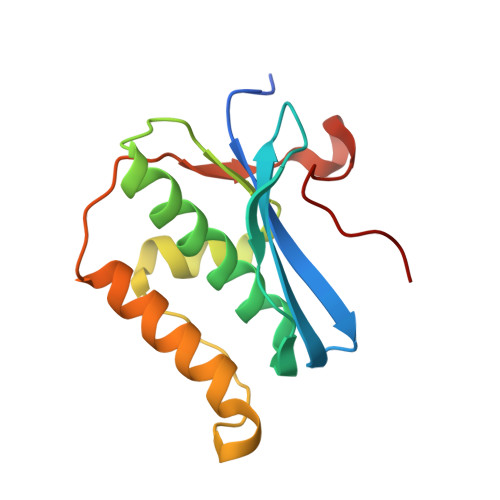
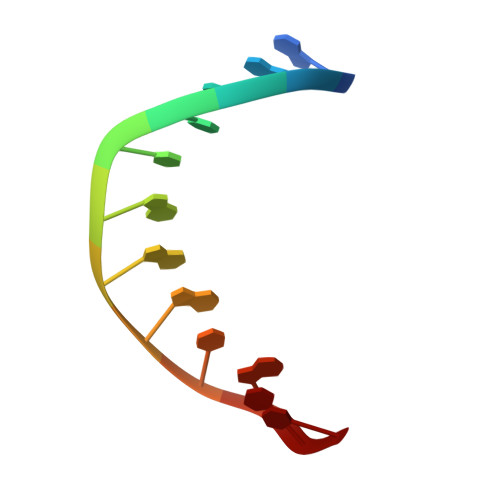
Crystal Structures of RNase H Bound to an RNA/DNA Hybrid: Substrate Specificity and Metal-Dependent Catalysis.
Nowotny, M., Gaidamakov, S.A., Crouch, R.J., Yang, W.(2005) Cell 121: 1005-1016
- PubMed: 15989951 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2005.04.024
- Primary Citation Related Structures:
1ZBF, 1ZBI, 1ZBL - PubMed Abstract:
RNase H belongs to a nucleotidyl-transferase superfamily, which includes transposase, retroviral integrase, Holliday junction resolvase, and RISC nuclease Argonaute. We report the crystal structures of RNase H complexed with an RNA/DNA hybrid and a mechanism for substrate recognition and two-metal-ion-dependent catalysis. RNase H specifically recognizes the A form RNA strand and the B form DNA strand. Structure comparisons lead us to predict the catalytic residues of Argonaute and conclude that two-metal-ion catalysis is a general feature of the superfamily. In nucleases, the two metal ions are asymmetrically coordinated and have distinct roles in activating the nucleophile and stabilizing the transition state. In transposases, they are symmetrically coordinated and exchange roles to alternately activate a water and a 3'-OH for successive strand cleavage and transfer by a ping-pong mechanism.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: