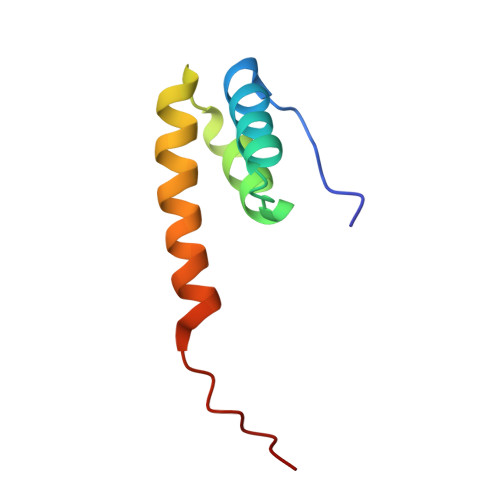
Structure of the functional domain of phi29 replication organizer: insights into oligomerization and dna binding
Asensio, J.L., Albert, A., Munoz-Espin, D., Gonzalez, C., Hermoso, J., Villar, L., Jimenez-Barbero, J., Salas, M., Meijer, W.J.J.(2005) J Biological Chem 280: 20730-20739
- PubMed: 15772069 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M501687200
- Primary Citation Related Structures:
1ZAE, 2BNK - PubMed Abstract:
The Bacillus subtilis phage phi29-encoded membrane protein p16.7 is one of the few proteins involved in prokaryotic membrane-associated DNA replication that has been characterized at a functional and biochemical level. In this work we have determined both the solution and crystal structures of its dimeric functional domain, p16.7C. Although the secondary structure of p16.7C is remarkably similar to that of the DNA binding homeodomain, present in proteins belonging to a large family of eukaryotic transcription factors, the tertiary structures of p16.7C and homeodomains are fundamentally different. In fact, p16.7C defines a novel dimeric six-helical fold. We also show that p16.7C can form multimers in solution and that this feature is a key factor for efficient DNA binding. Moreover, a combination of NMR and x-ray approaches, combined with functional analyses of mutants, revealed that multimerization of p16.7C dimers is mediated by a large protein surface that is characterized by a striking self-complementarity. Finally, the structural analyses of the p16.7C dimer and oligomers provide important clues about how protein multimerization and DNA binding are coupled.
- Departamento de Química Orgánica Biológica, Instituto de Química Orgánica General, Consejo Superior de Investigaciones Científicas, Madrid, Spain.
Organizational Affiliation: