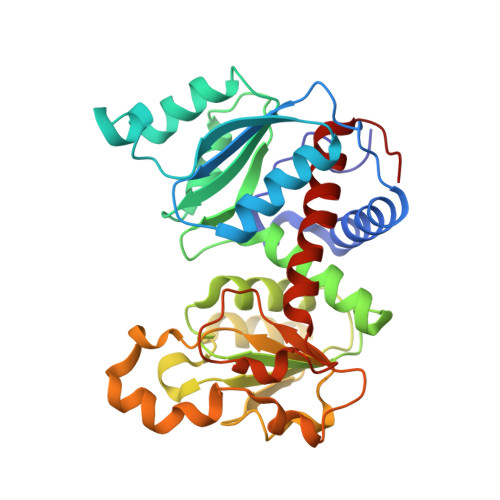
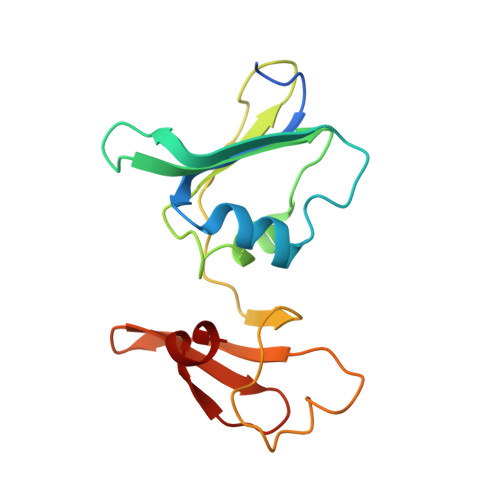
Structural basis for ordered substrate binding and cooperativity in aspartate transcarbamoylase
Wang, J., Stieglitz, K.A., Cardia, J.P., Kantrowitz, E.R.(2005) Proc Natl Acad Sci U S A 102: 8881-8886
- PubMed: 15951418 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0503742102
- Primary Citation Related Structures:
1ZA1, 1ZA2 - PubMed Abstract:
X-ray structures of aspartate transcarbamoylase in the absence and presence of the first substrate carbamoyl phosphate are reported. These two structures in conjunction with in silico docking experiments provide snapshots of critical events in the function of the enzyme. The ordered substrate binding, observed experimentally, can now be structurally explained by a conformational change induced upon the binding of carbamoyl phosphate. This induced fit dramatically alters the electrostatics of the active site, creating a binding pocket for aspartate. Upon aspartate binding, a further change in electrostatics causes a second induced fit, the domain closure. This domain closure acts as a clamp that both facilitates catalysis by approximation and also initiates the global conformational change that manifests homotropic cooperativity.
- Department of Chemistry, Boston College, Merkert Chemistry Center, Chestnut Hill, MA 02467, USA.
Organizational Affiliation: