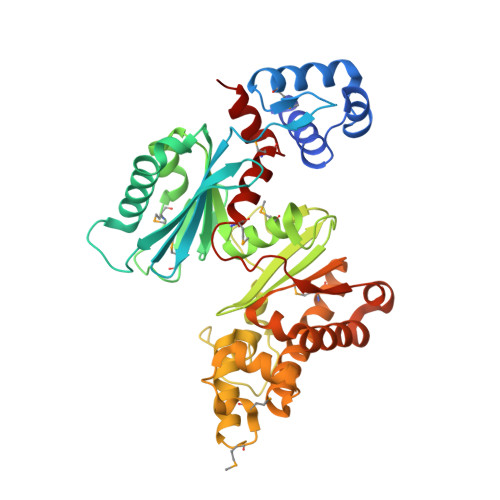
The crystal structure of Mlc, a global regulator of sugar metabolism in Escherichia coli
Schiefner, A., Gerber, K., Seitz, S., Welte, W., Diederichs, K., Boos, W.(2005) J Biological Chem 280: 29073-29079
- PubMed: 15929984 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M504215200
- Primary Citation Related Structures:
1Z6R - PubMed Abstract:
Mlc from Escherichia coli is a transcriptional repressor controlling the expression of a number of genes encoding enzymes of the phosphotransferase system (PTS), including ptsG and manXYZ, the specific enzyme II for glucose and mannose PTS transporters. In addition, Mlc controls the transcription of malT, the gene of the global activator of the mal regulon. The inactivation of Mlc as a repressor is mediated by binding to an actively transporting PtsG (EIICB(Glc)). Here we report the crystal structure of Mlc at 2.7 A resolution representing the first described structure of an ROK (repressors, open reading frames, and kinases) family protein. Mlc forms stable dimers thus explaining its binding affinity to palindromic operator sites. The N-terminal helix-turn-helix domain of Mlc is stabilized by the amphipathic C-terminal helix implicated earlier in EIICB(Glc) binding. Furthermore, the structure revealed a metal-binding site within the cysteine-rich ROK consensus motif that coordinates a structurally important zinc ion. A strongly reduced repressor activity was observed when two of the zinc-coordinating cysteine residues were exchanged against serine or alanine, demonstrating the role of zinc in Mlc-mediated repressor function. The structures of a putative fructokinase from Bacillus subtilis, the glucokinase from Escherichia coli, and a glucomannokinase from Arthrobacter sp. showed high structural homology to the ROK family part of Mlc.
- Department of Biology, University of Konstanz, Universitätsstrasse 10, 78457 Konstanz, Germany. andre.schiefner@uni-konstanz.de
Organizational Affiliation: