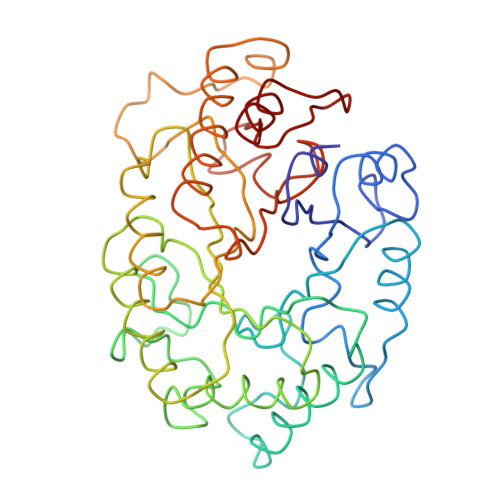
Structure of the catalytic core of the family F xylanase from Pseudomonas fluorescens and identification of the xylopentaose-binding sites.
Harris, G.W., Jenkins, J.A., Connerton, I., Cummings, N., Lo Leggio, L., Scott, M., Hazlewood, G.P., Laurie, J.I., Gilbert, H.J., Pickersgill, R.W.(1994) Structure 2: 1107-1116
- PubMed: 7881909 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(94)00112-x
- Primary Citation Related Structures:
1XYS - PubMed Abstract:
Sequence alignment suggests that xylanases evolved from two ancestral proteins and therefore can be grouped into two families, designated F and G. Family F enzymes show no sequence similarity with any known structure and their architecture is unknown. Studies of an inactive enzyme-substrate complex will help to elucidate the structural basis of binding and catalysis in the family F xylanases. We have therefore determined the crystal structure of the catalytic domain of a family F enzyme, Pseudomonas fluorescens subsp. cellulosa xylanase A, at 2.5 A resolution and a crystallographic R-factor of 0.20. The structure was solved using an engineered catalytic core in which the nucleophilic glutamate was replaced by a cysteine. As expected, this yielded both high-quality mercurial derivatives and an inactive enzyme which enabled the preparation of the inactive enzyme-substrate complex in the crystal. We show that family F xylanases are eight-fold alpha/beta-barrels (TIM barrels) with two active-site glutamates, one of which is the nucleophile and the other the acid-base. Xylopentaose binds to five subsites A-E with the cleaved bond between subsites D and E. Ca2+ binding, remote from the active-site glutamates, stabilizes the structure and may be involved in the binding of extended substrates. The architecture of P. fluorescens subsp. cellulosa has been determined crystallographically to be a commonly occurring enzyme fold, the eight-fold alpha/beta-barrel. Xylopentaose binds across the carboxy-terminal end of the alpha/beta-barrel in an active-site cleft which contains the two catalytic glutamates.
- Protein Engineering Department, Institute of Food Research, Reading Laboratory, UK.
Organizational Affiliation: