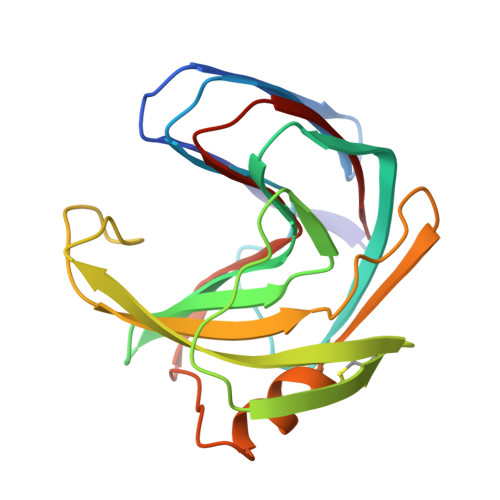
Thermostabilization of the Bacillus circulans xylanase by the introduction of disulfide bonds.
Wakarchuk, W.W., Sung, W.L., Campbell, R.L., Cunningham, A., Watson, D.C., Yaguchi, M.(1994) Protein Eng 7: 1379-1386
- PubMed: 7700870 Search on PubMed
- DOI: https://doi.org/10.1093/protein/7.11.1379
- Primary Citation Related Structures:
1XNC - PubMed Abstract:
The thermostability of the 20 396 Da Bacillus circulans xylanase was increased by the introduction of both intra- and intermolecular disulfide bridges by site-directed mutagenesis. Based on the 3-D structure of the enzyme, sites were chosen where favourable geometry for a bridge existed; in one case, to obtain favourable geometry additional mutations around the cysteine sites were designed by computer modelling. The disulfide bonds introduced into the xylanase were mostly buried and, in the absence of protein denaturants, relatively insensitive to reduction by dithiothreitol. The mutant proteins were examined for residual enzymatic activity after various thermal treatments, and were assayed for enzymatic activity at elevated temperatures to assess their productivity. We have examined one of these mutants by X-ray crystallography. All of the disulfide bond designs tested increased the thermostability of the B. circulans xylanase, but not all enhanced the activity of the enzyme at elevated temperatures.
- Institute for Biological Sciences, National Research Council of Canada, Ottawa, Ontario.
Organizational Affiliation: